Last universal common ancestor
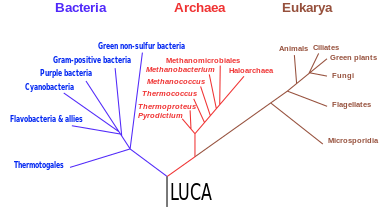
The last universal common ancestor (LUCA) is the hypothesized common ancestral cell from which the three domains of life, the Bacteria, the Archaea, and the Eukarya originated. The cell had a lipid bilayer; it possessed the genetic code and ribosomes which translated from DNA or RNA to proteins. The LUCA probably existed at latest 3.6 billion years ago, and possibly as early as 4.3 billion years ago[2] or earlier. The nature of this point or stage of divergence remains a topic of research.
All earlier forms of life preceding this divergence and all extant organisms are generally thought to share common ancestry. On the basis of a formal statistical test, this theory of a universal common ancestry (UCA) is supported versus competing multiple-ancestry hypotheses. The first universal common ancestor (FUCA) is a hypothetical non-cellular ancestor to LUCA and other now-extinct sister lineages.
Whether the genesis of viruses falls before or after the LUCA–as well as the diversity of extant viruses and their hosts–remains a subject of investigation.
While no fossil evidence of the LUCA exists, the detailed biochemical similarity of all current life (divided into the three domains) makes its existence widely accepted by biochemists. Its characteristics can be inferred from shared features of modern genomes. These genes describe a complex life form with many co-adapted features, including transcription and translation mechanisms to convert information from DNA to mRNA to proteins.
Historical background
[edit]
A phylogenetic tree directly portrays the idea of evolution by descent from a single ancestor.[3] An early tree of life was sketched by Jean-Baptiste Lamarck in his Philosophie zoologique in 1809.[4][5] Charles Darwin more famously proposed the theory of universal common descent through an evolutionary process in his book On the Origin of Species in 1859: "Therefore I should infer from analogy that probably all the organic beings which have ever lived on this earth have descended from some one primordial form, into which life was first breathed."[6] The last sentence of the book begins with a restatement of the hypothesis:
There is grandeur in this view of life, with its several powers, having been originally breathed into a few forms or into one ...
— [6]
The term "last universal common ancestor" or "LUCA" was first used in the 1990s for such a primordial organism.[7][8][9]
Inferring LUCA's features
[edit]An anaerobic thermophile
[edit]In 2016, Madeline C. Weiss and colleagues genetically analyzed 6.1 million protein-coding genes and 286,514 protein clusters from sequenced prokaryotic genomes representing many phylogenetic trees, and identified 355 protein clusters that were probably common to the LUCA. The results of their analysis are highly specific, though debated. They depict LUCA as "anaerobic, CO2-fixing, H2-dependent with a Wood–Ljungdahl pathway (the reductive acetyl-coenzyme A pathway), N2-fixing and thermophilic. LUCA's biochemistry was replete with FeS clusters and radical reaction mechanisms."[11] The cofactors also reveal "dependence upon transition metals, flavins, S-adenosyl methionine, coenzyme A, ferredoxin, molybdopterin, corrins and selenium. Its genetic code required nucleoside modifications and S-adenosylmethionine-dependent methylations."[11] They show that methanogenic clostridia were basal, near the root of the phylogenetic tree, in the 355 protein lineages examined, and that the LUCA may therefore have inhabited an anaerobic hydrothermal vent setting in a geochemically active environment rich in H2, CO2, and iron, where ocean water interacted with hot magma beneath the ocean floor.[11] It is even inferred that LUCA also grew from H2 and CO2 via the reverse incomplete Krebs cycle.[12] Other metabolic pathways inferred in LUCA are the pentose phosphate pathway, glycolysis, and gluconeogenesis.[13] Even if phylogenetic evidence may point to a hydrothermal vent environment for a thermophilic LUCA, this does not constitute evidence that the origin of life took place at a hydrothermal vent since mass extinctions may have removed previously existing branches of life.[14]
While the gross anatomy of the LUCA can be reconstructed only with much uncertainty, its biochemical mechanisms can be described in some detail, based on the "universal" properties currently shared by all independently living organisms on Earth.[15]

The LUCA certainly had genes and a genetic code.[10] Its genetic material was most likely DNA,[16] so that it lived after the RNA world.[a][19] The DNA was kept double-stranded by an enzyme, DNA polymerase, which recognises the structure and directionality of DNA.[20] The integrity of the DNA was maintained by a group of repair enzymes including DNA topoisomerase.[21] If the genetic code was based on dual-stranded DNA, it was expressed by copying the information to single-stranded RNA. The RNA was produced by a DNA-dependent RNA polymerase using nucleotides similar to those of DNA.[16] It had multiple DNA-binding proteins, such as histone-fold proteins.[22] The genetic code was expressed into proteins. These were assembled from 20 free amino acids by translation of a messenger RNA via a mechanism of ribosomes, transfer RNAs, and a group of related proteins.[16]
LUCA was likely capable of sexual interaction in the sense that adaptive gene functions were present that promoted the transfer of DNA between individuals of the population to facilitate genetic recombination. Homologous gene products that promote genetic recombination are present in bacteria, archaea and eukaryota, such as the RecA protein in bacteria, the RadA protein in archaea, and the Rad51 and Dmc1 proteins in eukaryota.[23]
The functionality of LUCA as well as evidence for the early evolution membrane-dependent biological systems together suggest that LUCA had cellularity and cell membranes.[24] As for the cell's gross structure, it contained a water-based cytoplasm effectively enclosed by a lipid bilayer membrane; it was capable of reproducing by cell division.[25] It tended to exclude sodium and concentrate potassium by means of specific ion transporters (or ion pumps). The cell multiplied by duplicating all its contents followed by cellular division. The cell used chemiosmosis to produce energy. It also reduced CO2 and oxidized H2 (methanogenesis or acetogenesis) via acetyl-thioesters.[26][27]
By phylogenetic bracketing, analysis of the presumed LUCA's offspring groups, LUCA appears to have been a small, single-celled organism. It likely had a ring-shaped coil of DNA floating freely within the cell. Morphologically, it would likely not have stood out within a mixed population of small modern-day bacteria. The originator of the three-domain system, Carl Woese, stated that in its genetic machinery, the LUCA would have been a "simpler, more rudimentary entity than the individual ancestors that spawned the three [domains] (and their descendants)".[1]
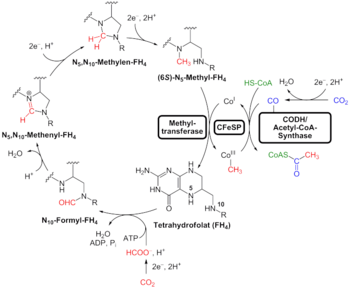
An alternative to the search for "universal" traits is to use genome analysis to identify phylogenetically ancient genes. This gives a picture of a LUCA that could live in a geochemically harsh environment and is like modern prokaryotes. Analysis of biochemical pathways implies the same sort of chemistry as does phylogenetic analysis. Weiss and colleagues write that "Experiments ... demonstrate that ... acetyl-CoA pathway [chemicals used in anaerobic respiration] formate, methanol, acetyl moieties, and even pyruvate arise spontaneously ... from CO2, native metals, and water", a combination present in hydrothermal vents.[28]
An experiment shows that Zn2+, Cr3+, and Fe can promote 6 of the 11 reactions of an ancient anabolic pathway called the reverse Krebs cycle in acidic conditions which implies that LUCA might have inhabited either hydrothermal vents or acidic metal-rich hydrothermal fields.[29]
Because both bacteria and archaea have differences in the structure of phospholipids and cell wall, ion pumping, most proteins involved in DNA replication, and glycolysis, it is inferred that LUCA had a permeable membrane without an ion pump. The emergence of Na+/H+ antiporters likely lead to the evolution of impermeable membranes present in eukaryotes, archaea, and bacteria. It is stated that "The late and independent evolution of glycolysis but not gluconeogenesis is entirely consistent with LUCA being powered by natural proton gradients across leaky membranes. Several discordant traits are likely to be linked to the late evolution of cell membranes, notably the cell wall, whose synthesis depends on the membrane and DNA replication".[30] Although LUCA likely had DNA, it is unknown if it could replicate DNA and is suggested to "might just have been a chemically stable repository for RNA-based replication".[31] It is likely that the permeable membrane of LUCA was composed of archaeal lipids (isoprenoids) and bacterial lipids (fatty acids). Isoprenoids would have enhanced stabilization of LUCA's membrane in the surrounding extreme habitat. Nick Lane and coauthors state that "The advantages and disadvantages of incorporating isoprenoids into cell membranes in different microenvironments may have driven membrane divergence, with the later biosynthesis of phospholipids giving rise to the unique G1P and G3P headgroups of archaea and bacteria respectively. If so, the properties conferred by membrane isoprenoids place the lipid divide as early as the origin of life".[32]
A 2024 study suggests that LUCA's genome was similar in size to that of modern prokaryotes, coding for some 2,600 proteins; that it respired anaerobically, and was an acetogen; and that it had an early CAS-based anti-viral immune system.[33]
Alternative interpretations
[edit]Some other researchers have challenged Weiss et al.'s 2016 conclusions. Sarah Berkemer and Shawn McGlynn argue that Weiss et al. undersampled the families of proteins, so that the phylogenetic trees were not complete and failed to describe the evolution of proteins correctly. There are two risks in attempting to attribute LUCA's environment from near-universal gene distribution (as in Weiss et al. 2016). On the one hand, it risks misattributing convergence or horizontal gene transfer events to vertical descent; on the other hand, it risks misattributing potential LUCA gene families as horizontal gene transfer events. A phylogenomic and geochemical analysis of a set of proteins that probably traced to the LUCA show that it had K+-dependent GTPases and the ionic composition and concentration of its intracellular fluid was seemingly high K+/Na+ ratio, NH+
4, Fe2+, CO2+, Ni2+, Mg2+, Mn2+, Zn2+, pyrophosphate, and PO3−
4 which would imply a terrestrial hot spring habitat. It possibly had a phosphate-based metabolism. Further, these proteins were unrelated to autotrophy (the ability of an organism to create its own organic matter), suggesting that the LUCA had a heterotrophic lifestyle (consuming organic matter) and that its growth was dependent on organic matter produced by the physical environment.[34] Nick Lane argues that Na+/H+ antiporters could readily explain the low concentration of Na+ in the LUCA and its descendants.
The presence of the energy-handling enzymes CODH/acetyl-coenzyme A synthase in LUCA could be compatible not only with being an autotroph but also with life as a mixotroph or heterotroph.[35] Weiss et al. 2018 reply that no enzyme defines a trophic lifestyle, and that heterotrophs evolved from autotrophs.[36]
Evidence that LUCA was mesophilic
[edit]Several lines of evidence now suggest that LUCA was non-thermophilic.
The content of G + C nucleotide pairs (compared to the occurrence of A + T pairs) can indicate an organism's thermal optimum as they are more thermally stable due to an additional hydrogen bond. As a result they occur more frequently in the rRNA of thermophiles; however this is not seen in LUCA's reconstructed rRNA.[37][38][14]
The identification of thermophilic genes in the LUCA has been criticized,[39] as they may instead represent genes that evolved later in archaea or bacteria, then migrated between these via horizontal gene transfer, as in Woese's 1998 hypothesis.[40] LUCA could have been a mesophile that fixed CO2 and relied on H2, and lived close to hydrothermal vents.[41]
Further evidence that LUCA was mesophilic comes from the amino acid composition of its proteins. The abundance of I, V, Y, W, R, E, and L amino acids (denoted IVYWREL) in an organism's proteins is correlated with its optimal growth temperature.[42] According to phylogentic analysis, the IVYWREL content of LUCA's proteins suggests its ideal temperature was below 50°C.[14]
Finally, evidence that bacteria and archaea both independently underwent phases of increased and subsequently decreased thermo-tolerance suggests a dramatic post-LUCA climate shift that affected both populations and would explain the seeming genetic pervasiveness of thermo-tolerant genetics.[43]
Age
[edit]Studies from 2000 to 2018 have suggested an increasingly ancient time for the LUCA. In 2000, estimates of the LUCA's age ranged from 3.5 to 3.8 billion years ago in the Paleoarchean,[44] a few hundred million years before the earliest fossil evidence of life, for which candidates range in age from 3.48 to 4.28 billion years ago.[45][46][47][48][49] This placed the origin of the first forms of life shortly after the Late Heavy Bombardment which was thought to have repeatedly sterilized Earth's surface. However, a 2018 study by Holly Betts and colleagues applied a molecular clock model to the genomic and fossil record (102 species, 29 common protein-coding genes, mostly ribosomal), concluding that LUCA preceded the Late Heavy Bombardment (making the LUCA over 3.9 billion years ago). They assumed that there was no sterilizing event after the Moon-forming event, which they dated as 4.520 billion years ago, and concluded that the most likely date for LUCA was within 50 million years of that.[50] A 2022 study suggested an age of around 3.6-4.2 billion years for the LUCA.[51] A 2024 study suggested that the LUCA lived around 4.2 billion years ago (with a confidence interval of 4.09–4.33 billion years ago).[33]
Root of the tree of life
[edit]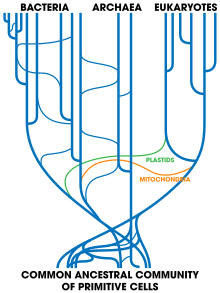
In 1990, a novel concept of the tree of life was presented, dividing the living world into three stems, classified as the domains Bacteria, Archaea, Eukarya.[1][53][54][55] It is the first tree founded exclusively on molecular phylogenetics, and which includes the evolution of microorganisms. It has been called a "universal phylogenetic tree in rooted form".[1] This tree and its rooting became the subject of debate.[53][b]
In the meantime, numerous modifications of this tree, mainly concerning the role and importance of horizontal gene transfer for its rooting and early ramifications have been suggested (e.g.[57][52]). Since heredity occurs both vertically and horizontally, the tree of life may have been more weblike or netlike in its early phase and more treelike when it grew three-stemmed.[52] Presumably horizontal gene transfer has decreased with growing cell stability.[58]
A modified version of the tree, based on several molecular studies, has its root between a monophyletic domain Bacteria and a clade formed by Archaea and Eukaryota.[57] A small minority of studies place the root in the domain bacteria, in the phylum Bacillota,[59] or state that the phylum Chloroflexota (formerly Chloroflexi) is basal to a clade with Archaea and Eukaryotes and the rest of bacteria (as proposed by Thomas Cavalier-Smith).[60] Metagenomic analyses recover a two-domain system with the domains Archaea and Bacteria; in this view of the tree of life, Eukaryotes are derived from Archaea.[61][62][63] With the later gene pool of LUCA's descendants, sharing a common framework of the AT/GC rule and the standard twenty amino acids, horizontal gene transfer would have become feasible and could have been common.[64]
The nature of LUCA remains disputed. In 1994, on the basis of primordial metabolism (sensu Wächtershäuser), Otto Kandler proposed a successive divergence of the three domains of life[1] from a multiphenotypical population of pre-cells, reached by gradual evolutionary improvements (cellularization).[65][66][67] These phenotypically diverse pre-cells were metabolising, self-reproducing entities exhibiting frequent mutual exchange of genetic information. Thus, in this scenario there was no "first cell". It may explain the unity and, at the same time, the partition into three lines (the three domains) of life. Kandler's pre-cell theory is supported by Wächtershäuser.[68][69] In 1998, Carl Woese, based on the RNA world concept, proposed that no individual organism could be considered a LUCA, and that the genetic heritage of all modern organisms derived through horizontal gene transfer among an ancient community of organisms.[70] Other authors concur that there was a "complex collective genome"[71] at the time of the LUCA, and that horizontal gene transfer was important in the evolution of later groups;[71] Nicolas Glansdorff states that LUCA "was in a metabolically and morphologically heterogeneous community, constantly shuffling around genetic material" and "remained an evolutionary entity, though loosely defined and constantly changing, as long as this promiscuity lasted."[72]
The theory of a universal common ancestry of life is widely accepted. In 2010, based on "the vast array of molecular sequences now available from all domains of life,"[73] D. L. Theobald published a "formal test" of universal common ancestry (UCA). This deals with the common descent of all extant terrestrial organisms, each being a genealogical descendant of a single species from the distant past. His formal test favoured the existence of a universal common ancestry over a wide class of alternative hypotheses that included horizontal gene transfer. Basic biochemical principles imply that all organisms do have a common ancestry.[74]
A proposed, earlier, non-cellular ancestor to LUCA is the First universal common ancestor (FUCA).[75][76] FUCA would therefore be the ancestor to every modern cell as well as ancient, now-extinct cellular lineages not descendant of LUCA. FUCA is assumed to have had other descendants than LUCA, none of which have modern descendants. Some genes of these ancient now-extinct cell lineages are thought to have been horizontally transferred into the genome of early descendants of LUCA.[64]
LUCA and viruses
[edit]The origin of viruses remains disputed. Since viruses need host cells for their replication, it is likely that they emerged after the formation of cells. Viruses may even have multiple origins and different types of viruses may have evolved independently over the history of life.[55] There are different hypotheses for the origins of viruses, for instance an early viral origin from the RNA world or a later viral origin from selfish DNA.[55]
Based on how viruses are currently distributed across the bacteria and archaea, the LUCA is suspected of having been prey to multiple viruses, ancestral to those that now have those two domains as their hosts.[77] Furthermore, extensive virus evolution seems to have preceded the LUCA, since the jelly-roll structure of capsid proteins is shared by RNA and DNA viruses across all three domains of life.[78][79] LUCA's viruses were probably mainly dsDNA viruses in the groups called Duplodnaviria and Varidnaviria. Two other single-stranded DNA virus groups within the Monodnaviria, the Microviridae and the Tubulavirales, likely infected the last bacterial common ancestor. The last archaeal common ancestor was probably host to spindle-shaped viruses. All of these could well have affected the LUCA, in which case each must since have been lost in the host domain where it is no longer extant. By contrast, RNA viruses do not appear to have been important parasites of LUCA, even though straightforward thinking might have envisaged viruses as beginning with RNA viruses directly derived from an RNA world. Instead, by the time the LUCA lived, RNA viruses had probably already been out-competed by DNA viruses.[77]
LUCA might have been the ancestor to some viruses, as it might have had at least two descendants: LUCELLA, the Last Universal Cellular Ancestor, the ancestor to all cells, and the archaic virocell ancestor, the ancestor to large-to-medium-sized DNA viruses.[80] Viruses might have evolved before LUCA but after the First universal common ancestor (FUCA), according to the reduction hypothesis, where giant viruses evolved from primordial cells that became parasitic.[64]
See also
[edit]- Abiogenesis – Life arising from non-living matter
- Cellularization – Scientific theory to explain the origin and formation of cells
- Chemoton – Abstract model for the fundamental unit of life
- Darwinian threshold – Period during the evolution of the first cells
- Last eukaryotic common ancestor – Process of forming the first eukaryotic cell
- Mitochondrial Eve – Matrilineal most recent common ancestor of all living humans
- Pre-cell – Hypothetical life before complete cells
- Proto-metabolism
- Timeline of the evolutionary history of life
- Urmetazoan – Hypothetical last common ancestor of all animals
- Y-chromosomal Adam – Patrilineal most recent common ancestor of all living humans
Notes
[edit]- ^ Other studies propose that LUCA may have been defined wholly through RNA,[17] consisted of a RNA-DNA hybrid genome, or possessed a retrovirus-like genetic cycle with DNA serving as a stable genetic repository.[18]
- ^ One debate dealt with a former cladistic hypothesis: The tree could not be ascribed a root in the usual algorithmic way, because that would require an outgroup for reference. In the case of the universal tree, no outgroup would exist. The cladistic method was used "to root the purple bacteria, for example. But establishing a root for the universal tree of life, the branching order among the primary urkingdoms, was another matter entirely."[56]
References
[edit]- ^ a b c d e Woese, C.R.; Kandler, O.; Wheelis, M.L. (June 1990). "Towards a natural system of organisms: proposal for the domains Archaea, Bacteria, and Eucarya". PNAS. 87 (12): 4576–4579. Bibcode:1990PNAS...87.4576W. doi:10.1073/pnas.87.12.4576. PMC 54159. PMID 2112744.
- ^ Moody, Edmund; Álvarez-Carretero, Sandra; Mahendrarajah, Tara (12 July 2024). "The nature of the last universal common ancestor and its impact on the early Earth system". Nat. Ecol. Evol. doi:10.1038/s41559-024-02461-1. PMC 11383801. Retrieved 26 August 2024.
- ^ Gregory, T. Ryan (2008). "Understanding evolutionary trees". Evolution: Education and Outreach. 1 (2): 121–137. doi:10.1007/s12052-008-0035-x. S2CID 15488906.
- ^ Lamarck, Jean Baptiste Pierre Antoine de Monet de (1994) [1809]. Philosophie zoologique (PDF). Paris. p. 737.
{{cite book}}: CS1 maint: location missing publisher (link) - ^ Noble, Denis (1 July 2020). "Editorial: Charles Darwin, Jean-Baptiste Lamarck, and 21st century arguments on the fundamentals of biology". Progress in Biophysics and Molecular Biology. 153: 1–4. doi:10.1016/j.pbiomolbio.2020.02.005. PMID 32092299. S2CID 211475380. Archived from the original on 1 March 2022. Retrieved 23 December 2022.
- ^ a b Darwin, Charles (1859). The Origin of Species by Means of Natural Selection. John Murray. pp. 484, 490. Archived from the original on 8 October 2022. Retrieved 8 October 2022.
- ^ Wikham, Gene Stephen (March 1995). The molecular phylogenetic analysis of naturally occurring hyperthermophilic microbial communities (PhD thesis). Indiana University. p. 4. ProQuest 304192982
- ^ Forterre, Patrick (1997). "Archaea: What can we learn from their sequences?". Current Opinion in Genetics & Development. 7 (6): 764–770. doi:10.1016/s0959-437x(97)80038-x. PMID 9468785.
- ^ Koonin, Eugene V.; Galperin, Michael Y. (2003). Sequence - Evolution - Function: Computational approaches in comparative genomics. Boston, MA: Kluwer. p. 252. ISBN 978-1-4757-3783-7. OCLC 55642057.
- ^ a b c Weiss, Madeline C.; Preiner, Martina; Xavier, Joana C.; Zimorski, Verena; Martin, William F. (16 August 2018). "The last universal common ancestor between ancient Earth chemistry and the onset of genetics". PLOS Genetics. 14 (8): e1007518. doi:10.1371/journal.pgen.1007518. PMC 6095482. PMID 30114187. S2CID 52019935.
- ^ a b c Weiss, Madeline C.; Sousa, F. L.; Mrnjavac, N.; et al. (2016). "The physiology and habitat of the last universal common ancestor" (PDF). Nature Microbiology. 1 (9): 16116. doi:10.1038/nmicrobiol.2016.116. PMID 27562259. S2CID 2997255. Archived (PDF) from the original on 18 April 2022. Retrieved 10 October 2022.
- ^ Harrison, Stuart A.; Palmeira, Raquel Nunes; Halpern, Aaron; Lane, Nick (1 November 2022). "A biophysical basis for the emergence of the genetic code in protocells". Biochimica et Biophysica Acta (BBA) - Bioenergetics. 1863 (8): 148597. doi:10.1016/j.bbabio.2022.148597. PMID 35868450.
- ^ Harrison, Stuart A.; Lane, Nick (12 December 2018). "Life as a guide to prebiotic nucleotide synthesis". Nature Communications. 9 (1): 5176. Bibcode:2018NatCo...9.5176H. doi:10.1038/s41467-018-07220-y. ISSN 2041-1723. PMC 6289992. PMID 30538225.
- ^ a b c Cantine, Marjorie D.; Fournier, Gregory P. (6 July 2017). "Environmental Adaptation from the Origin of Life to the Last Universal Common Ancestor". Origins of Life and Evolution of Biospheres. 48 (1): 35–54. doi:10.1007/s11084-017-9542-5. hdl:1721.1/114219. PMID 28685374. S2CID 254888920. Archived from the original on 23 February 2024. Retrieved 4 December 2023.
- ^ Wächtershäuser, Günter (1998). "Towards a Reconstruction of Ancestral Genomes by Gene Cluster Alignment". Systematic and Applied Microbiology. 21 (4): 473–474, IN1, 475–477. Bibcode:1998SyApM..21N1475W. doi:10.1016/S0723-2020(98)80058-1.
- ^ a b c Wächtershäuser, Günter (1998). "Towards a Reconstruction of Ancestral Genomes by Gene Cluster Alignment". Systematic and Applied Microbiology. 21 (4): 473–474, IN1, 475–477. Bibcode:1998SyApM..21N1475W. doi:10.1016/S0723-2020(98)80058-1.
- ^ Marshall, Michael. "Life began with a planetary mega-organism". New Scientist. Archived from the original on 25 July 2016. Retrieved 31 July 2016.
- ^ Koonin, Eugene V.; Martin, William (1 December 2005). "On the origin of genomes and cells within inorganic compartments". Trends in Genetics. 21 (12): 647–654. doi:10.1016/j.tig.2005.09.006. PMC 7172762. PMID 16223546.
- ^ Garwood, Russell J. (2012). "Patterns In Palaeontology: The first 3 billion years of evolution". Palaeontology Online. 2 (11): 1–14. Archived from the original on 26 June 2015. Retrieved 25 June 2015.
- ^ Koonin, Eugene V.; Krupovic, M.; Ishino, S.; Ishino, Y. (2020). "The replication machinery of LUCA: common origin of DNA replication and transcription". BMC Biology. 18 (1): 61. doi:10.1186/s12915-020-00800-9. PMC 7281927. PMID 32517760.
- ^ Ahmad, Muzammil; Xu, Dongyi; Wang, Weidong (23 May 2017). "Type IA topoisomerases can be "magicians" for both DNA and RNA in all domains of life". RNA Biology. 14 (7): 854–864. doi:10.1080/15476286.2017.1330741. PMC 5546716. PMID 28534707.
- ^ Lupas, Andrei N.; Alva, Vikram (2018). "Histones predate the split between bacteria and archaea". Bioinformatics. 35 (14): 2349–2353. doi:10.1093/bioinformatics/bty1000. PMID 30520969.
- ^ Bernstein, H., Bernstein, C. (2017). Sexual Communication in Archaea, the Precursor to Eukaryotic Meiosis. In: Witzany, G. (eds) Biocommunication of Archaea. Springer, Cham. https://doi.org/10.1007/978-3-319-65536-9_7 Archived 23 February 2024 at the Wayback Machine
- ^ Gogarten, Johann Peter; Taiz, Lincoln (1992). "Evolution of proton pumping ATPases: Rooting the tree of life". Photosynthesis Research. 33 (2): 137–146. Bibcode:1992PhoRe..33..137G. doi:10.1007/bf00039176. ISSN 0166-8595. PMID 24408574. S2CID 20013957. Archived from the original on 23 February 2024. Retrieved 4 December 2023.
- ^ Wächtershäuser, Günter (1998). "Towards a Reconstruction of Ancestral Genomes by Gene Cluster Alignment". Systematic and Applied Microbiology. 21 (4): 473–474, IN1, 475–477. Bibcode:1998SyApM..21N1475W. doi:10.1016/S0723-2020(98)80058-1.
- ^ Martin, W.; Russell, M. J. (October 2007). "On the origin of biochemistry at an alkaline hydrothermal vent". Philosophical Transactions of the Royal Society of London. Series B, Biological Sciences. 362 (1486): 1887–1925. doi:10.1098/rstb.2006.1881. PMC 2442388. PMID 17255002.
- ^ Lane, Nick; Allen, J. F.; Martin, William (April 2010). "How did LUCA make a living? Chemiosmosis in the origin of life". BioEssays. 32 (4): 271–280. doi:10.1002/bies.200900131. PMID 20108228.
- ^ Weiss, Madeline C.; Preiner, Martina; Xavier, Joana C.; Zimorski, Verena; Martin, William F. (16 August 2018). "The last universal common ancestor between ancient Earth chemistry and the onset of genetics". PLOS Genetics. 14 (8): e1007518. doi:10.1371/journal.pgen.1007518. PMC 6095482. PMID 30114187. S2CID 52019935.
- ^ Muchowska, Kamila B.; Varma, Sreejith J.; Chevallot-Beroux, Elodie; Lethuillier-Karl, Lucas; Li, Guang; Moran, Joseph (2 October 2017). "Metals promote sequences of the reverse Krebs cycle". Nature Ecology & Evolution. 1 (11): 1716–1721. Bibcode:2017NatEE...1.1716M. doi:10.1038/s41559-017-0311-7. ISSN 2397-334X. PMC 5659384. PMID 28970480.
- ^ Sojo, Víctor; Pomiankowski, Andrew; Lane, Nick (12 August 2014). "A Bioenergetic Basis for Membrane Divergence in Archaea and Bacteria". PLOS Biology. 12 (8): e1001926. doi:10.1371/journal.pbio.1001926. PMC 4130499. PMID 25116890.
- ^ Weiss, Madeline C.; Preiner, Martina; Xavier, Joana C.; Zimorski, Verena; Martin, William F. (16 August 2018). "The last universal common ancestor between ancient Earth chemistry and the onset of genetics". PLOS Genetics. 14 (8): e1007518. doi:10.1371/journal.pgen.1007518. PMC 6095482. PMID 30114187. S2CID 52019935.
- ^ Jordan, S. F.; Nee, E.; Lane, Nick (18 October 2019). "Isoprenoids enhance the stability of fatty acid membranes at the emergence of life potentially leading to an early lipid divide". Interface Focus. 9 (6). doi:10.1098/rsfs.2019.0067. PMC 6802135. PMID 31641436.
 This article incorporates text from this source, which is available under the CC BY 4.0 license.
This article incorporates text from this source, which is available under the CC BY 4.0 license.
- ^ a b Moody, Edmund R. R.; Álvarez-Carretero, Sandra; Mahendrarajah, Tara A.; Clark, James W.; Betts, Holly C.; Dombrowski, Nina; Szánthó, Lénárd L.; Boyle, Richard A.; Daines, Stuart; Chen, Xi; Lane, Nick; Yang, Ziheng; Shields, Graham A.; Szöllősi, Gergely J.; Spang, Anja (12 July 2024). "The nature of the last universal common ancestor and its impact on the early Earth system". Nature Ecology & Evolution: 1–13. doi:10.1038/s41559-024-02461-1. ISSN 2397-334X. PMC 11383801.
- ^ Mulkidjanian, Armen Y.; Bychkov, Andrew Yu; Dibrova, Daria V.; Galperin, Michael Y.; Koonin, Eugene V. (2012). "Origin of first cells at terrestrial, anoxic geothermal fields". Proceedings of the National Academy of Sciences. 109 (14): E821-30. Bibcode:2012PNAS..109E.821M. doi:10.1073/pnas.1117774109. PMC 3325685. PMID 22331915.
- ^ Adam, Panagiotis S.; Borrel, Guillaume; Gribaldo, Simonetta (6 February 2018). "Evolutionary history of carbon monoxide dehydrogenase/acetyl-CoA synthase, one of the oldest enzymatic complexes". PNAS. 115 (6): E1166–E1173. Bibcode:2018PNAS..115E1166A. doi:10.1073/pnas.1716667115. PMC 5819426. PMID 29358391.
- ^ Weiss, Madeline C.; Preiner, Martina; Xavier, Joana C.; Zimorski, Verena; Martin, William F. (16 August 2018). "The last universal common ancestor between ancient Earth chemistry and the onset of genetics". PLOS Genetics. 14 (8): e1007518. doi:10.1371/journal.pgen.1007518. PMC 6095482. PMID 30114187. S2CID 52019935.
- ^ Galtier, Nicolas; Tourasse, Nicolas; Gouy, Manolo (8 January 1999). "A Nonhyperthermophilic Common Ancestor to Extant Life Forms". Science. 283 (5399): 220–221. doi:10.1126/science.283.5399.220. ISSN 0036-8075. PMID 9880254. Archived from the original on 23 February 2024. Retrieved 4 December 2023.
- ^ Groussin, Mathieu; Boussau, Bastien; Charles, Sandrine; Blanquart, Samuel; Gouy, Manolo (23 October 2013). "The molecular signal for the adaptation to cold temperature during early life on Earth". Biology Letters. 9 (5): 20130608. doi:10.1098/rsbl.2013.0608. PMC 3971708. PMID 24046876.
- ^ Gogarten, Johann Peter; Deamer, David (2016). "Is LUCA a thermophilic progenote?". Nature Microbiology. 1 (12): 16229. doi:10.1038/nmicrobiol.2016.229. PMID 27886195. S2CID 205428194. Archived from the original on 3 April 2020. Retrieved 25 June 2019.
- ^ Woese, Carl (June 1998). "The universal ancestor". PNAS. 95 (12): 6854–6859. Bibcode:1998PNAS...95.6854W. doi:10.1073/pnas.95.12.6854. PMC 22660. PMID 9618502.
- ^ Camprubí, E.; de Leeuw, J. W.; House, C. H.; Raulin, F.; Russell, M. J.; Spang, A.; Tirumalai, M. R.; Westall, F. (12 December 2019). "The Emergence of Life". Space Science Reviews. 215 (8): 56. Bibcode:2019SSRv..215...56C. doi:10.1007/s11214-019-0624-8. ISSN 1572-9672.
- ^ Zeldovich, Konstantin B; Berezovsky, Igor N; Shakhnovich, Eugene I (2007). "Protein and DNA Sequence Determinants of Thermophilic Adaptation". PLOS Computational Biology. 3 (1): e5. arXiv:q-bio/0607004. Bibcode:2007PLSCB...3....5Z. doi:10.1371/journal.pcbi.0030005. ISSN 1553-7358. PMC 1769408. PMID 17222055.
- ^ Boussau, Bastien; Blanquart, Samuel; Necsulea, Anamaria; Lartillot, Nicolas; Gouy, Manolo (26 November 2008). "Parallel adaptations to high temperatures in the Archaean eon". Nature. 456 (7224): 942–945. Bibcode:2008Natur.456..942B. doi:10.1038/nature07393. PMID 19037246. S2CID 4348746. Archived from the original on 23 February 2024. Retrieved 4 December 2023.
- ^ Doolittle, W. F. (February 2000). "Uprooting the tree of life". Scientific American. 282 (2): 90–95. Bibcode:2000SciAm.282b..90D. doi:10.1038/scientificamerican0200-90. JSTOR 26058605. PMID 10710791.
- ^ Noffke, N.; Christian, D.; Wacey, D.; Hazen, R. M. (December 2013). "Microbially induced sedimentary structures recording an ancient ecosystem in the ca. 3.48 billion-year-old Dresser Formation, Pilbara, Western Australia". Astrobiology. 13 (12): 1103–1124. Bibcode:2013AsBio..13.1103N. doi:10.1089/ast.2013.1030. PMC 3870916. PMID 24205812.
- ^ Ohtomo, Yoko; Kakegawa, Takeshi; Ishida, Akizumi; Nagase, Toshiro; Rosing, Minik T. (2013). "Evidence for biogenic graphite in early Archaean Isua metasedimentary rocks". Nature Geoscience. 7 (1): 25–28. Bibcode:2014NatGe...7...25O. doi:10.1038/ngeo2025.
- ^ Hassenkam, T.; Andersson, M. P.; Dalby, K. N.; et al. (2017). "Elements of Eoarchean life trapped in mineral inclusions". Nature. 548 (7665): 78–81. Bibcode:2017Natur.548...78H. doi:10.1038/nature23261. PMID 28738409. S2CID 205257931.
- ^ Bell, Elizabeth A.; Boehnke, Patrick; Harrison, T. Mark; Mao, Wendy L. (24 November 2015). "Potentially biogenic carbon preserved in a 4.1 billion-year-old zircon". PNAS. 112 (47): 14518–14521. Bibcode:2015PNAS..11214518B. doi:10.1073/pnas.1517557112. PMC 4664351. PMID 26483481.
- ^ Dodd, Matthew S.; Papineau, Dominic; Grenne, Tor; et al. (2 March 2017). "Evidence for early life in Earth's oldest hydrothermal vent precipitates" (PDF). Nature. 543 (7643): 60–64. Bibcode:2017Natur.543...60D. doi:10.1038/nature21377. PMID 28252057. S2CID 2420384. Archived (PDF) from the original on 23 July 2018. Retrieved 25 June 2019.
- ^ Betts, Holly C.; Puttick, Mark N.; Clark, James W.; Williams, Tom A.; Donoghue, Philip C. J.; Pisani, Davide (2018). "Integrated genomic and fossil evidence illuminates life's early evolution and eukaryote origin". Nature Ecology & Evolution. 2 (10): 1556–1562. Bibcode:2018NatEE...2.1556B. doi:10.1038/s41559-018-0644-x. PMC 6152910. PMID 30127539.
- ^ Moody, Edmund RR; Mahendrarajah, Tara A.; Dombrowski, Nina; Clark, James W.; Petitjean, Celine; Offre, Pierre; Szöllősi, Gergely J.; Spang, Anja; Williams, Tom A. (22 February 2022). "An estimate of the deepest branches of the tree of life from ancient vertically evolving genes". eLife. 11. doi:10.7554/eLife.66695. ISSN 2050-084X. PMC 8890751. PMID 35190025.
- ^ a b c Smets, Barth F.; Barkay, Tamar (September 2005). "Horizontal gene transfer: perspectives at a crossroads of scientific disciplines". Nature Reviews Microbiology. 3 (9): 675–678. doi:10.1038/nrmicro1253. PMID 16145755. S2CID 2265315.
- ^ a b Sapp, Jan A. (2009). The new foundations of evolution: on the tree of life. New York: Oxford University Press. pp. Chapter 19: 257ff (novel concept of the tree of life), Chapters 17-21 plus concluding remarks: 226-318 (discussion of the tree and its rooting), 286ff (LUCA). ISBN 978-0-199-73438-2. Archived from the original on 6 November 2023. Retrieved 21 November 2023.
- ^ Madigan, Michael T.; Martinko, John M.; Bender, Kelly S.; Buckley, Daniel H.; Stahl, David A. (2015). Brock Biology of Microorganisms (14 ed.). Boston: Pearson Education Limited. pp. 29, 374, 381. ISBN 978-1-292-01831-7.
- ^ a b c Madigan, Michael T.; Aiyer, Jennifer; Buckley, Daniel H.; Sattley, Matthew; Stahl, David A. (2022). Brock Biology of Microorganisms (16 ed.). Harlow: Pearson Education. pp. Unit 3, chapter 13: 431 (LUCA), 435 (tree of life), 428, 438, 439 (viruses). ISBN 978-1-292-40479-0.
- ^ Sapp 2009, p. 255.
- ^ a b Brown, J. R.; Doolittle, W. F. (1995). "Root of the Universal Tree of Life Based on Ancient Aminoacyl-tRNA Synthetase Gene Duplications". PNAS. 92 (7): 2441–2445. Bibcode:1995PNAS...92.2441B. doi:10.1073/pnas.92.7.2441. PMC 42233. PMID 7708661.
- ^ Harold, Franklin M. (2014). In Search of Cell History: The Evolution of Life's Building Blocks. Chicago, London: University of Chicago Press. ISBN 978-0-226-17428-0. Archived from the original on 31 October 2023. Retrieved 12 October 2023.
- ^ Valas, R. E.; Bourne, P. E. (2011). "The origin of a derived superkingdom: how a gram-positive bacterium crossed the desert to become an archaeon". Biology Direct. 6: 16. doi:10.1186/1745-6150-6-16. PMC 3056875. PMID 21356104.
- ^ Cavalier-Smith, Tom (2006). "Rooting the tree of life by transition analyses". Biology Direct. 1: 19. doi:10.1186/1745-6150-1-19. PMC 1586193. PMID 16834776.
- ^ Raymann, Kasie; Brochier-Armanet, Céline; Gribaldo, Simonetta (26 May 2015). "The two-domain tree of life is linked to a new root for the Archaea". Proceedings of the National Academy of Sciences. 112 (21): 6670–6675. Bibcode:2015PNAS..112.6670R. doi:10.1073/pnas.1420858112. ISSN 0027-8424. PMC 4450401. PMID 25964353.
- ^ Hug, Laura A.; Baker, Brett J.; Anantharaman, Karthik; et al. (11 April 2016). "A new view of the tree of life". Nature Microbiology. 1 (5): 16048. doi:10.1038/nmicrobiol.2016.48. ISSN 2058-5276. PMID 27572647. S2CID 3833474.
- ^ Williams, Tom A.; Foster, Peter G.; Cox, Cymon J.; Embley, T. Martin (11 December 2013). "An archaeal origin of eukaryotes supports only two primary domains of life". Nature. 504 (7479): 231–236. Bibcode:2013Natur.504..231W. doi:10.1038/nature12779. ISSN 1476-4687. PMID 24336283. S2CID 4461775. Archived from the original on 1 October 2022. Retrieved 23 September 2022.
- ^ a b c Harris, Hugh M. B.; Hill, Colin (2021). "A Place for Viruses on the Tree of Life". Frontiers in Microbiology. 11. doi:10.3389/fmicb.2020.604048. ISSN 1664-302X. PMC 7840587. PMID 33519747.
- ^ Kandler, Otto (1994). "The early diversification of life". In Stefan Bengtson (ed.). Early Life on Earth. Nobel Symposium 84. New York: Columbia University Press. pp. 152–160.
- ^ Kandler, Otto (1995). "Cell Wall Biochemistry in Archaea and its Phylogenetic Implications". Journal of Biological Physics. 20 (1–4): 165–169. doi:10.1007/BF00700433. S2CID 83906865.
- ^ Kandler, Otto (1998). "The early diversification of life and the origin of the three domains: A proposal". In Jürgen Wiegel; Michael W. W. Adams (eds.). Thermophiles: The keys to molecular evolution and the origin of life?. London: Taylor and Francis Ltd. pp. 19–31. ISBN 978-0-203-48420-3. Archived from the original on 25 February 2023. Retrieved 21 June 2023.
- ^ Wächtershäuser, Günter (2003). "From pre-cells to Eukarya – a tale of two lipids". Molecular Microbiology. 47 (1): 13–22. doi:10.1046/j.1365-2958.2003.03267.x. PMID 12492850. S2CID 37944519.
- ^ Wächtershäuser, Günter (October 2006). "From volcanic origins of chemoautotrophic life to Bacteria, Archaea and Eukarya". Philosophical Transactions of the Royal Society of London. Series B, Biological Sciences. 361 (1474): 1787–1808. doi:10.1098/rstb.2006.1904. PMC 1664677. PMID 17008219.
- ^ Woese, Carl (June 1998). "The universal ancestor". PNAS. 95 (12): 6854–6859. Bibcode:1998PNAS...95.6854W. doi:10.1073/pnas.95.12.6854. PMC 22660. PMID 9618502.
- ^ a b Egel, Richard (March 2012). "Primal Eukaryogenesis: On the Communal Nature of Precellular States, Ancestral to Modern Life". Life. 2 (1): 170–212. Bibcode:2012Life....2..170E. doi:10.3390/life2010170. PMC 4187143. PMID 25382122.
- ^ Glansdorff, Nicolas; Xu, Ying; Labedan, Bernard (9 July 2008). "The Last Universal Common Ancestor: emergence, constitution and genetic legacy of an elusive forerunner". Biology Direct. 3 (1): 29. doi:10.1186/1745-6150-3-29. PMC 2478661. PMID 18613974. S2CID 18250196.
- ^ Steel, M.; Penny, D. (May 2010). "Origins of life: Common ancestry put to the test". Nature. 465 (7295): 168–169. Bibcode:2010Natur.465..168S. doi:10.1038/465168a. PMID 20463725. S2CID 205055573.
- ^ Theobald, D. L. (May 2010). "A formal test of the theory of universal common ancestry". Nature. 465 (7295): 219–222. Bibcode:2010Natur.465..219T. doi:10.1038/nature09014. PMID 20463738. S2CID 4422345.
- ^ Prosdocimi, Francisco; José, Marco V.; de Farias, Sávio Torres (2019). "The First Universal Common Ancestor (FUCA) as the Earliest Ancestor of LUCA's (Last UCA) Lineage". In Pontarotti, Pierre (ed.). Evolution, Origin of Life, Concepts and Methods. Cham: Springer. pp. 43–54. doi:10.1007/978-3-030-30363-1_3. ISBN 978-3-030-30363-1. S2CID 199534387. Retrieved 2 November 2023.
- ^ Prosdocimi, Francisco; José, Marco V.; de Farias, Sávio Torres (2019), "The First Universal Common Ancestor (FUCA) as the Earliest Ancestor of LUCA's (Last UCA) Lineage", in Pontarotti, Pierre (ed.), Evolution, Origin of Life, Concepts and Methods, Cham: Springer, pp. 43–54, doi:10.1007/978-3-030-30363-1_3, ISBN 978-3-030-30363-1, S2CID 199534387, archived from the original on 23 February 2024, retrieved 2 November 2023
- ^ a b Krupovic, M.; Dolja, V. V.; Koonin, Eugene V. (2020). "The LUCA and its complex virome" (PDF). Nature Reviews Microbiology. 18 (11): 661–670. doi:10.1038/s41579-020-0408-x. PMID 32665595. S2CID 220516514. Archived (PDF) from the original on 21 October 2022. Retrieved 15 August 2021.
- ^ Forterre, Patrick; Prangishvili, David (2009). "The origin of viruses". Research in Microbiology. 160 (7): 466–472. doi:10.1016/j.resmic.2009.07.008. PMID 19647075. S2CID 2767388.
- ^ Durzyńska, Julia; Goździcka-Józefiak, Anna (16 October 2015). "Viruses and cells intertwined since the dawn of evolution". Virology Journal. 12 (1): 169. doi:10.1186/s12985-015-0400-7. PMC 4609113. PMID 26475454.
- ^ Nasir, Arshan; Kim, Kyung Mo; Caetano-Anollés, Gustavo (1 September 2012). "Viral evolution". Mobile Genetic Elements. 2 (5): 247–252. doi:10.4161/mge.22797. ISSN 2159-2543. PMC 3575434. PMID 23550145.
Further reading
[edit]- Lane, Nick (2016) [2015]. The Vital Question. London: Profile Books. ISBN 978-1781250372.


