Haplogroup D-M174
| Haplogroup D-M174 | |
|---|---|
 | |
| Possible time of origin | 50,000[1] - 60,000[2] years BP 65,200 [95% CI 62,100 <-> 68,300] ybp[3] |
| Coalescence age | 46,300 [95% CI 43,500 <-> 49,100] ybp[3] |
| Possible place of origin | Asia[2][4][5] (possibly Central Asia[6] or Southeast Asia[2]) |
| Ancestor | D(D-CTS3946) |
| Descendants | D-Z27276(D1a1) D-M55(D1a2) D-Y34637(D1a3) |
| Defining mutations | M174, IMS-JST021355, PAGES00003 |
Haplogroup D1 or D-M174 is a subclade of Haplogroup D-CTS3946. This haplogroup is found primarily in East Asia and the Andaman Islands, though it is also found regularly with low frequency in Central Asia and Southeast Asia.
Origins
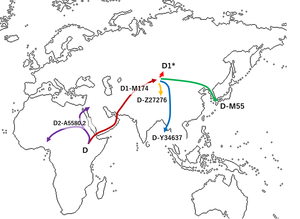
Haplogroup D-M174 is believed to have originated in Asia some 60,000 years before present, It has been suggested by Shi et al. (2008) to have a Southeast Asian (rather than North or Central Asian) origin due to its rarity in North and Central Asia.[2][4] While haplogroup D-M174 along with haplogroup E contains the distinctive YAP polymorphism (which indicates their common ancestry), no haplogroup D-M174 chromosomes have been found anywhere outside of Asia.[4]
Several studies (Hammer et al. 2006, Shinoda 2008, Matsumoto 2009, Cabrera et al. 2018) suggest that the paternal haplogroup D-M174 originated somewhere in Central Asia. According to Hammer et al., haplogroup D-M174 originated between Tibet and the Altai mountains. He suggests that there were multiple waves into Eastern Eurasia.[7][why?]
A 2017 study by Mondal et al. finds that the Riang people (a Tibeto-Burmese population) and Andamanese have their nearest related lineages in East Asia. The Jarawa and Onge shared this D1a3 lineages with each other within the last ~7000 years, but had diverged from the Japanese D1a2 lineage ~53000 years ago". They further suggest that: ”This strongly suggests that haplogroup D does not indicate a separate ancestry for Andamanese populations. Rather, haplogroup D was part of the standing variation carried by the OOA expansion, and later lost from most of the populations except in Andaman and partially in Japan and Tibet”.[8]
A study (Haber et al. 2019) found a proposed haplogroup named "D0" in three Nigerian samples. In part because of the likely deep-rooting of haplogroup "D0", as well as recently calculated early divergence times for it and its parent haplogroup, the authors suggest an African origin for D0 and its parent haplogroup DE. The "D0" haplogroup is outside M174, but shares 7 SNPs with it that E lacks.[9] "D0" has also been named "D2", and former D (D-M174) has now also been termed "D1" due to this discovery.
Three other samples of D2 were also found in West Asia (also in 2019): two in Saudi Arabia and another one in Syria. The sample found in the Syrian is to date the most basal sample of D2. The recent evidence (as also proposed by Haber et al.) suggests that D2 is a highly divergent haplogroup close to the DE split but on the D branch and lacking the M174 mutation possessed by other D lineages.[10]
Overview
It is found today at high frequency among populations in Tibet, northern Myanmar, Qinghai, the Japanese archipelago, and the Andaman Islands, though curiously not as much in the rest of India. The Ainu of Japan and various Tibeto-Burmese people (such as the Tripuri people) are notable for possessing almost exclusively Haplogroup D-M174 chromosomes. Haplogroup D-M174 chromosomes are also found at low to moderate frequencies among the Bai, Dai, Han, Hui, Manchu, Miao, Tujia, Xibe, Yao, and Zhuang of China and among several minority populations of Sichuan and Yunnan that speak Tibeto-Burman languages and reside in close proximity to the Tibetans, such as the Jingpo, Jino, Mosuo, Naxi, Pumi, Qiang, and Yi.[11]
Haplogroup D is also found in populations of China proper and Korea, but with much lower frequency than in populations of Tibet and Japan. A study published in 2011 found D-M174 in 2.49% (43/1729) of Han Chinese males, with frequencies of this haplogroup tending to be higher than average toward the north and toward the west of the country (5/56 = 8.9% D-M174 Shaanxi Han, 13/221 = 5.9% D-M174 Gansu Han, 6/136 = 4.4% D-M174 Yunnan Han, 1/27 = 3.7% D-M174 Guangxi Han, 2/61 = 3.3% D-M174 Hunan Han, 2/62 = 3.2% D-M174 Sichuan Han).[12] In another study of Han Chinese Y-DNA published in 2011, Haplogroup D-M174 was observed in 1.94% (7/361) of a sample of unrelated Han Chinese male volunteers at Fudan University in Shanghai, with the origins of most of the volunteers being traced back to East China (Jiangsu, Zhejiang, Shanghai, and Anhui).[13] In Korea, Haplogroup D-M174 has been observed in 3.8% (5/133) of a sample from Daejeon,[14] 3/85 = 3.5% of a sample from Seoul,[15] 3.3% (3/90) of a sample from Jeolla,[16] 2.4% (2/84) of a sample from Gyeongsang,[16] 2.3% (13/573) of another sample from Seoul,[14] 1.4% (1/72) of a sample from Chungcheong,[16] 1.1% (1/87) of a sample from Jeju,[16] and 0.9% (1/110) of a third sample from Seoul-Gyeonggi.[16] In other studies, haplogroup D-M174 has been observed in 6.7% (3/45)[17] and 4.0% (3/75)[18] of samples from Korea without any further specification of the area of sampling. Little high-resolution data regarding the phylogenetic position of Han Chinese and Korean members of Y-DNA Haplogroup D has been published, but the available data suggests that most Han Chinese members of Haplogroup D should belong to clades found frequently among Tibetans (and especially the D-M15 clade, also found among speakers of some Lolo-Burmese and Hmong-Mien languages), whereas most Korean members of Haplogroup D should belong to the D-M55 clade, which is found frequently among Ainu, Ryukyuan, and Japanese people.[18][16][3]
Haplogroup D Y-DNA has been found (albeit with low frequency) among modern populations of the Eurasian steppe, such as Southern Altaians (6/96 = 6.3% D-M174(xM15),[19] 6/120 = 5.0% D-P47[20]), Kazakhs (1/54 = 1.9% D-M174,[17] 6/1294 = 0.5% D[21]), Nogais (4/76 = 5.3% D-M174 Kara Nogai,[22] 1/87 = 1.1% D-M174 Kuban Nogai[22]), Khalkhas (1/24 = 4.2% D-M174,[17] 3/85 = 3.5% D-M174,[15] 2/149 D-M15 + 2/149 D-P47 = 4/149 = 2.7% D-M174 total[18]), Zakhchin (2/60 = 3.3% D-M174[15]), Uriankhai (1/60 = 1.7% D-M174[15]), and Kalmyks (5/426 = 1.2% D-M174[23]). It also has been found among linguistically similar (Turkic- or Mongolic-speaking) modern populations of the desert and oasis belt south of the steppe, such as Yugurs, Bao’an, Monguors, Uyghurs, and Uzbeks. In commercial testing, members have been found as far west as Romania in Europe and Iraq in Western Asia.[24]
Unlike haplogroup C-M217, Haplogroup D-M174 is not found in the New World; it is not present in any modern Native American (North, Central or South) populations. While it is possible that it traveled to the New World like Haplogroup C-M217, those lineages apparently became extinct.
Haplogroup D-M174 is also remarkable for its rather extreme geographic differentiation, with a distinct subset of Haplogroup D-M174 chromosomes being found exclusively in each of the populations that contains a large percentage of individuals whose Y-chromosomes belong to Haplogroup D-M174: Haplogroup D-M15 among the Tibetans (as well as among other East Asian and Southeast Asian populations that display low frequencies of Haplogroup D-M174 Y-chromosomes), Haplogroup D-M55 among the various populations of the Japanese Archipelago and with low frequency among Koreans, Haplogroup D-P99 among the inhabitants of Tibet and some other parts of central Eurasia (e.g. Mongolia[25] and the Altai[18][19][20]). D-M174* without positive-tested of subclades D-M15 or D-M55 is found at high frequencies among Andaman Islanders and recently Andamanese subclade is found to be D-Y34637 (D1a3).[26] Another type (or types) of paragroup D-M174* without tested positive subclades of D-M15,D-P47,or D-M55 is found at a very low frequency among the Turkic and Mongolic populations of Central Asia, amounting to no more than 1% in total. This apparently ancient diversification of Haplogroup D-M174 suggests that it may perhaps be better characterized as a "super-haplogroup" or "macro-haplogroup."
In one study, the frequency of Haplogroup D-M174 without tested positive subclades found among Thais was 10%.[2] Su et al. (2000) found DE-YAP/DYS287(xM15) in 11.1% (5/45) of a set of three samples from Thailand (including 20% (4/20) North Thai, 20% (1/5) So, and 0% (0/20) Northeast Thai) and in 16.7% (1/6) of a sample from Guam.[27] Meanwhile, the authors found D-M15 in 15% of a pair of samples of Yao (including 30% (3/10) Yao Jinxiu and 0% (0/10) Yao Nandan), 14.3% (2/14) of a sample of Yi, 3.8% (1/26) of a sample of Cambodians, and 3.6% (1/28) of a sample of Zhuang.[27] Dong et al. (2002) found DE-YAP Y-chromosomes in 12.5% (2/16) of a sample of Jingpo from Luxi City, Yunnan, 10.0% (2/20) of a sample of Dai from Luxi City, Yunnan, and 1.82% (1/55) of a sample of Nu from Gongshan and Fugong counties of Yunnan.[28]
Distribution
The Haplogroup D-M174 Y-chromosomes that are found among Tibeto-Burman populations as well as people of the Japanese Archipelago (haplogroup D1a3, D1a2 and D1a1). D-M55 (D1a2) are particularly distinctive, bearing a complex of at least five individual mutations along an internal branch of the Haplogroup D-M174 phylogeny, thus distinguishing them clearly from the other Haplogroup D-M174 chromosomes that are found among the Tibetans and Andaman Islanders and providing evidence that Y-chromosome Haplogroup D-M55 was the modal haplogroup in the ancestral population that developed the prehistoric Jōmon culture in the Japanese islands.
It is suggested that the majority of D-M174 Y-chromosome carriers migrated from Central Asia to East Asia. One group migrated into the Andaman Islands, thus forming or helping to form the Andamanese people. Another group stayed in modern Tibet and southern China (today Tibeto-Burman peoples) and another group migrated to Japan, possibly via the Korean Peninsula (pre-Jōmon people).[29][30]
Subclades
D-Z27276 (D1a1)
Haplogroup D-Z27276 is the common ancestor of D-M15 and D-P99.
D-M15 (D1a1a)
D-M15 was first reported to have been found in a sample from Cambodia and Laos (1/18 = 5.6%) and in a sample from Japan (1/23 = 4.3%) in a preliminary worldwide survey of Y-DNA variation in extant human populations.[31]
Subsequently, Y-DNA that belongs to Haplogroup D-M15 has been found frequently among Tibeto-Burman-speaking populations of Southwestern China (including approximately 23% of Qiang,[2][32][33] approximately 12.5% of Tibetans,[2] and approximately 9% of Yi[2][34]) and among Yao people inhabiting northeastern Guangxi (6/31 = 19.4% Lowland Yao, 5/41 = 12.2% Native Mien, 3/41 = 7.3% Lowland Kimmun)[35] with a moderate distribution throughout Central Asia, East Asia, and continental Southeast Asia (Indochina).[2]
A study published in 2011 has found D-M15 in 7.8% (4/51) of a sample of Hmong Daw and in 3.4% (1/29) of a sample of Xinhmul from northern Laos.[35]
D-P47 (D1a1b1)
Found with high frequency among Pumi,[2] Naxi,[2] and Tibetans,[36][2] with a moderate distribution in Central Asia.[2] According to one study, Tibetans have a frequency of about 41.31% of haplogroup D-P47.[37] Haplogroup D-P47 has been found in 9% of Xi'an Han.[38]
D-M55 (D1a2)
Previously known as D-M55, D-M64.1/Page44.1 (D1a2) is found with high frequency among Ainu,[39] and at medium frequency among Japanese,[40] and Ryukyuans.[40]
Kim et al. (2011) found Haplogroup D-M55 in 2.0% (1/51) of a sample of Beijing Han and in 1.6% (8/506) of a pool of samples from South Korea, including 3.3% (3/90) from the Jeolla region, 2.4% (2/84) from the Gyeongsang region, 1.4% (1/72) from the Chungcheong region, 1.1% (1/87) from the Jeju region, 0.9% (1/110) from the Seoul-Gyeonggi region, and 0% (0/63) from the Gangwon region.[16] Hammer et al. (2006) found Haplogroup D-P37.1 in 4.0% (3/75) of a sample from South Korea.[18]
Low levels of D-M116.1 (a subclade of D-M55) among males in present-day Timor (0.2% of males),[41] It's found in 9.5% of males from Micronesia(Hammer et al. 2006,[42][18] is believed to reflect recent admixture from Japan. That is, D-M116.1 (D1a2a) is generally believed to be a primary subclade of D-M64.1 (D1a2), possibly as a result of the Japanese military occupation of South East Asia during World War II.
According to Mitsuru Sakitani, Haplogroup D1 arrived from Central Asia to northern Kyushu via the Altai Mountains and the Korean Peninsula more than 40,000 years before present, and Haplogroup D-M55 (D1a2) was born in Japanese archipelago.[43]
Recently it was confirmed that the Japanese branch of haplogroup D-M55 is distinct and isolated from other D-branches since more than 53,000 years. The split between D1a probably happened in Central Asia, while some others suggest an instant split during the origin of haplogroup D itself, as the Japanese branch has five unique mutations not found in any other D-branch.[44]
(D*)Paragroup D1a3 (D34637)
D1a3 is found at high frequencies among Andaman Islanders[2] (especially Onge(23/23 = 100%), Jarawa(4/4 = 100%).[45][26]
D1a3* is found in some Tibetan minority tribes in Northeast India (among whom rates vary from zero to 65%).[5][46][47][48]
D-M175*
The basal D-M174* (xM15,P47,M55) has been found in approximately 5% of Altaians.[18] Kharkov et al. have found haplogroup D*(xD-M15) in 6.3% (6/96) of a pool of samples of Southern Altaians from three different localities, particularly in Kulada (5/46 = 10.9%) and Kosh-Agach (1/7 = 14%), though they have not tested for any marker of the subclade D-M55 or D-P99. Kharkov et al. also have reported finding haplogroup DE-M1(xD-M174) Y-DNA in one Southern Altaian individual from Beshpeltir (1/43 = 2.3%).[19]
Phylogenetics
Phylogenetic history
Prior to 2002, there were in academic literature at least seven naming systems for the Y-Chromosome Phylogenetic tree. This led to considerable confusion. In 2002, the major research groups came together and formed the Y-Chromosome Consortium (YCC). They published a joint paper that created a single new tree that all agreed to use. Latter, a group of citizen scientists with an interest in population genetics and genetic genealogy formed a working group to create an amateur tree aiming at being above all timely. The table below brings together all of these works at the point of the landmark 2002 YCC Tree. This allows a researcher reviewing older published literature to quickly move between nomenclatures.
| YCC 2002/2008 (Shorthand) | (α) | (β) | (γ) | (δ) | (ε) | (ζ) | (η) | YCC 2002 (Longhand) | YCC 2005 (Longhand) | YCC 2008 (Longhand) | YCC 2010r (Longhand) | ISOGG 2006 | ISOGG 2007 | ISOGG 2008 | ISOGG 2009 | ISOGG 2010 | ISOGG 2011 | ISOGG 2012 |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| D-M174 | * | * | * | * | * | * | * | * | D | D | D | D | D | D | D | D | D | D |
| D-M15 | 4 | IV | 3G | 12 | Eu5 | H3 | B | D1 | D1 | D1 | D1 | D1 | D1 | D1 | D1 | D1 | D1 | D1 |
| D-M55 | * | * | * | * | * | * | * | * | D2 | D2 | D2 | D2 | D2 | D2 | D2 | D2 | D2 | D2 |
| D-P12 | 4 | IV | 3G | 11 | Eu5 | H2 | B | D2a | D2a | D2a1a1 | D2a1a1 | D2 | D2 | D2a1a1 | D2a1a1 | D2a1a1 | removed | removed |
| D-M116.1 | 4 | IV | 3G | 11 | Eu5 | H2 | B | D2b* | D2a | D2a | D2a | D2a | D2a | D2a | D2a | D2a | removed | removed |
| D-M125 | 4 | IV | 3G | 11 | Eu5 | H2 | B | D2b1 | D2a1 | D2a1 | D2a1 | D2a1 | D2a1 | D2a1 | D2a1 | D2a1 | D2a1 | D2a1 |
| D-M151 | 4 | IV | 3G | 11 | Eu5 | H2 | B | D2b2 | D2a1 | D2a2 | D2a2 | D2a2 | D2a2 | D2a2 | D2a2 | D2a2 | D2a2 | D2a2 |
Research publications
The following research teams per their publications were represented in the creation of the YCC tree.
Phylogenetic trees
By ISOGG tree(Version: 14.151).[49]
- DE (YAP)
- D (CTS3946)
- D1 (M174/Page30, IMS-JST021355)
- D1a (CTS11577)
- D1a1 (F6251/Z27276)
- D1a1a (M15) Mainland China, Tibet
- D1a1b (P99) Mainland China, Tibet, Mongol, Central Asia
- D1a2 (M64.1/Page44.1, M55) Japan(Yamato people、Ryukyuan people、Ainu people)
- D1a3 (Y34637) Andaman Islands(Onge people、Jarawa people)
- D1a1 (F6251/Z27276)
- D1b (L1378) Philippines[50]
- D1a (CTS11577)
- D2 (A5580.2) Nigeria, Saudi Arabia, Syria
- D1 (M174/Page30, IMS-JST021355)
- D (CTS3946)
See also
Genetics
- Genetic genealogy
- Haplogroup
- Haplotype
- Human Y-chromosome DNA haplogroup
- Molecular phylogeny
- Paragroup
- Subclade
- Y-chromosome haplogroups in populations of the world
- Y-DNA haplogroups by ethnic group
- Y-DNA haplogroups in populations of East and Southeast Asia
- Y-DNA haplogroups in populations of Oceania
Y-DNA D-M174 subclades
Y-DNA backbone tree
References
- ^ "Y-DNA Haplogroup D-M174 and its Subclades - 2017".
- ^ a b c d e f g h i j k l m n Shi H, Zhong H, Peng Y, Dong YL, Qi XB, Zhang F, Liu LF, Tan SJ, Ma RZ, Xiao CJ, Wells RS, Jin L, Su B (October 2008). "Y chromosome evidence of earliest modern human settlement in East Asia and multiple origins of Tibetan and Japanese populations". BMC Biology. 6: 45. doi:10.1186/1741-7007-6-45. PMC 2605740. PMID 18959782.
{{cite journal}}: CS1 maint: unflagged free DOI (link) - ^ a b c YFull Haplogroup YTree v6.02 at 02 April 2018. Accessed July 7, 2018.
- ^ a b c Karafet TM, Mendez FL, Meilerman MB, Underhill PA, Zegura SL, Hammer MF (May 2008). "New binary polymorphisms reshape and increase resolution of the human Y chromosomal haplogroup tree". Genome Research. 18 (5): 830–8. doi:10.1101/gr.7172008. PMC 2336805. PMID 18385274.
- ^ a b Su B, Xiao C, Deka R, Seielstad MT, Kangwanpong D, Xiao J, Lu D, Underhill P, Cavalli-Sforza L, Chakraborty R, Jin L (December 2000). "Y chromosome haplotypes reveal prehistorical migrations to the Himalayas". Human Genetics. 107 (6): 582–90. doi:10.1007/s004390000406. PMID 11153912.
- ^ Gayden, Tenzin; Cadenas, Alicia M.; Regueiro, Maria; Singh, Nanda B.; Zhivotovsky, Lev A.; Underhill, Peter A.; Cavalli-Sforza, Luigi L.; Herrera, Rene J. (May 2007). "The Himalayas as a Directional Barrier to Gene Flow". The American Journal of Human Genetics. 80 (5): 884–894. doi:10.1086/516757. PMC 1852741. PMID 17436243.
- ^ Matsumoto H (February 2009). "The origin of the Japanese race based on genetic markers of immunoglobulin G". Proceedings of the Japan Academy. Series B, Physical and Biological Sciences. 85 (2): 69–82. Bibcode:2009PJAB...85...69M. doi:10.2183/pjab.85.69. PMC 3524296. PMID 19212099.
- ^ Mondal, Mayukh; Bergström, Anders; Xue, Yali; Calafell, Francesc; Laayouni, Hafid; Casals, Ferran; Majumder, Partha P.; Tyler-Smith, Chris; Bertranpetit, Jaume (2017-05-01). "Y-chromosomal sequences of diverse Indian populations and the ancestry of the Andamanese". Human Genetics. 136 (5): 499–510. doi:10.1007/s00439-017-1800-0. hdl:10230/34399. ISSN 1432-1203. PMID 28444560.
In contrast, the Riang (Tibeto-Burman-speaking) and Andamanese have their nearest neighbour lineages in East Asia. The Jarawa and Onge shared haplogroup D lineages with each other within the last ~7000 years, but had diverged from Japanese haplogroup D Y-chromosomes ~53000 years ago, most likely by a split from a shared ancestral population.
- ^ Tyler-Smith, Chris; Xue, Yali; Thomas, Mark G.; Yang, Huanming; Arciero, Elena; Asan; Connell, Bruce A.; Jones, Abigail L.; Haber, Marc (2019-06-13). "A Rare Deep-Rooting D0 African Y-Chromosomal Haplogroup and Its Implications for the Expansion of Modern Humans out of Africa". Genetics. 212 (4): 1421–1428. doi:10.1534/genetics.119.302368. ISSN 0016-6731. PMC 6707464. PMID 31196864.
- ^ Estes, Roberta (2019-06-21). "Exciting New Y DNA Haplogroup D Discoveries!". DNAeXplained - Genetic Genealogy. Retrieved 2019-07-08.
- ^ Y染色体单倍群D在東亞的分布及其意義
- ^ Zhong H, Shi H, Qi XB, Duan ZY, Tan PP, Jin L, Su B, Ma RZ (2011). "Extended Y Chromosome Investigation Suggests Postglacial Migrations of Modern Humans into East Asia via the Northern Route". Mol. Biol. Evol. 28 (1): 717–727. doi:10.1093/molbev/msq247. PMID 20837606.
- ^ Yan S, Wang CC, Li H, Li SL, Jin L, et al. (The Genographic Consortium) (September 2011). "An updated tree of Y-chromosome Haplogroup O and revised phylogenetic positions of mutations P164 and PK4". European Journal of Human Genetics. 19 (9): 1013–5. doi:10.1038/ejhg.2011.64. PMC 3179364. PMID 21505448.
- ^ a b Park MJ, Lee HY, Yang WI, Shin KJ (July 2012). "Understanding the Y chromosome variation in Korea--relevance of combined haplogroup and haplotype analyses". International Journal of Legal Medicine. 126 (4): 589–99. doi:10.1007/s00414-012-0703-9. PMID 22569803.
- ^ a b c d Katoh T, Munkhbat B, Tounai K, Mano S, Ando H, Oyungerel G, Chae GT, Han H, Jia GJ, Tokunaga K, Munkhtuvshin N, Tamiya G, Inoko H (February 2005). "Genetic features of Mongolian ethnic groups revealed by Y-chromosomal analysis". Gene. 346: 63–70. doi:10.1016/j.gene.2004.10.023. PMID 15716011.
- ^ a b c d e f g Kim, Soon-Hee; Kim, Ki-Cheol; Shin, Dong-Jik; Jin, Han-Jun; Kwak, Kyoung-Don; Han, Myun-Soo; Song, Joon-Myong; Kim, Won; Kim, Wook (2011). "High frequencies of Y-chromosome haplogroup O2b-SRY465 lineages in Korea: a genetic perspective on the peopling of Korea". Investigative Genetics. 2 (1): 10. doi:10.1186/2041-2223-2-10. PMC 3087676. PMID 21463511.
{{cite journal}}: CS1 maint: unflagged free DOI (link) - ^ a b c Wells RS, Yuldasheva N, Ruzibakiev R, Underhill PA, Evseeva I, Blue-Smith J, Jin L, Su B, Pitchappan R, Shanmugalakshmi S, Balakrishnan K, Read M, Pearson NM, Zerjal T, Webster MT, Zholoshvili I, Jamarjashvili E, Gambarov S, Nikbin B, Dostiev A, Aknazarov O, Zalloua P, Tsoy I, Kitaev M, Mirrakhimov M, Chariev A, Bodmer WF (August 2001). "The Eurasian heartland: a continental perspective on Y-chromosome diversity". Proceedings of the National Academy of Sciences of the United States of America. 98 (18): 10244–9. Bibcode:2001PNAS...9810244W. doi:10.1073/pnas.171305098. PMC 56946. PMID 11526236.
- ^ a b c d e f g Hammer MF, Karafet TM, Park H, Omoto K, Harihara S, Stoneking M, Horai S (2006). "Dual origins of the Japanese: common ground for hunter-gatherer and farmer Y chromosomes". Journal of Human Genetics. 51 (1): 47–58. doi:10.1007/s10038-005-0322-0. PMID 16328082.
- ^ a b c Khar'kov VN, Stepanov VA, Medvedeva OF, Spiridonova MG, Voevoda MI, Tadinova VN, Puzyrev VP (May 2007). "[Gene pool differences between northern and southern Altaians inferred from the data on Y-chromosomal haplogroups]". Genetika (in Russian). 43 (5): 675–87. doi:10.1134/S1022795407050110. PMID 17633562.
- ^ a b Dulik MC, Zhadanov SI, Osipova LP, Askapuli A, Gau L, Gokcumen O, Rubinstein S, Schurr TG (February 2012). "Mitochondrial DNA and Y chromosome variation provides evidence for a recent common ancestry between Native Americans and Indigenous Altaians". American Journal of Human Genetics. 90 (2): 229–46. doi:10.1016/j.ajhg.2011.12.014. PMC 3276666. PMID 22281367.
- ^ E. E. Ashirbekov, D. M. Botbaev, A. M. Belkozhaev, A. O. Abayldaev, A. S. Neupokoeva, J. E. Mukhataev, B. Alzhanuly, D. A. Sharafutdinova, D. D. Mukushkina, M. B. Rakhymgozhin, A. K. Khanseitova, S. A. Limborska, and N. A. Aytkhozhina, "Distribution of Y-Chromosome Haplogroups of the Kazakh from the South Kazakhstan, Zhambyl, and Almaty Regions." Reports of the National Academy of Sciences of the Republic of Kazakhstan, ISSN 2224-5227, Volume 6, Number 316 (2017), 85 - 95.
- ^ a b Yunusbayev B, Metspalu M, Järve M, Kutuev I, Rootsi S, Metspalu E, Behar DM, Varendi K, Sahakyan H, Khusainova R, Yepiskoposyan L, Khusnutdinova EK, Underhill PA, Kivisild T, Villems R (January 2012). "The Caucasus as an asymmetric semipermeable barrier to ancient human migrations". Molecular Biology and Evolution. 29 (1): 359–65. doi:10.1093/molbev/msr221. PMID 21917723.
- ^ Malyarchuk B, Derenko M, Denisova G, Khoyt S, Woźniak M, Grzybowski T, Zakharov I (December 2013). "Y-chromosome diversity in the Kalmyks at the ethnical and tribal levels". Journal of Human Genetics. 58 (12): 804–11. doi:10.1038/jhg.2013.108. PMID 24132124.
- ^ Y-DNA D Haplogroup Project at Family Tree DNA
- ^ Di Cristofaro J, Pennarun E, Mazières S, Myres NM, Lin AA, Temori SA, Metspalu M, Metspalu E, Witzel M, King RJ, Underhill PA, Villems R, Chiaroni J (2013). "Afghan Hindu Kush: where Eurasian sub-continent gene flows converge". PLOS ONE. 8 (10): e76748. Bibcode:2013PLoSO...876748D. doi:10.1371/journal.pone.0076748. PMC 3799995. PMID 24204668.
{{cite journal}}: CS1 maint: unflagged free DOI (link) - ^ a b Y-Full
- ^ a b Su, Bing; Jin, Li; Underhill, Peter; Martinson, Jeremy; Saha, Nilmani; McGarvey, Stephen T.; Shriver, Mark D.; Chu, Jiayou; Oefner, Peter; Chakraborty, Ranajit; Deka, Ranjan (18 July 2000). "Polynesian origins: Insights from the Y chromosome". Proceedings of the National Academy of Sciences of the United States of America. 97 (15): 8225–8228. Bibcode:2000PNAS...97.8225S. doi:10.1073/pnas.97.15.8225. PMC 26928. PMID 10899994.
- ^ DONG Yongli, SHI Hong, LI Weixiang, YANG Jie, ZENG Weimin, LI Kaiyuan, and XIAO Chunjie, "Study of polymorphism at the YAP locus in seven Yunnan ethnic minority populations in the great gorge of the Salween River and downstream areas" (original title in Chinese: "怒江大峡谷及下游地区7个云南少数民族YAP位点的多态性研究"), Acta Anthropologica Sinica, Vol. 21, No. 3 (August, 2002). http://www.ivpp.cas.cn/cbw/rlxxb/xbwzxz/201203/t20120320_3512811.html
- ^ Hammer MF, Karafet TM, Park H, Omoto K, Harihara S, Stoneking M, Horai S (2006). "Dual origins of the Japanese: common ground for hunter-gatherer and farmer Y chromosomes". Journal of Human Genetics. 51 (1): 47–58. doi:10.1007/s10038-005-0322-0. PMID 16328082.
- ^ Shi H, Zhong H, Peng Y, Dong YL, Qi XB, Zhang F, Liu LF, Tan SJ, Ma RZ, Xiao CJ, Wells RS, Jin L, Su B (October 2008). "Y chromosome evidence of earliest modern human settlement in East Asia and multiple origins of Tibetan and Japanese populations". BMC Biology. 6 (1): 45. doi:10.1186/1741-7007-6-45. PMC 2605740. PMID 18959782.
{{cite journal}}: CS1 maint: unflagged free DOI (link) - ^ Underhill PA, Shen P, Lin AA, Jin L, Passarino G, Yang WH, Kauffman E, Bonné-Tamir B, Bertranpetit J, Francalacci P, Ibrahim M, Jenkins T, Kidd JR, Mehdi SQ, Seielstad MT, Wells RS, Piazza A, Davis RW, Feldman MW, Cavalli-Sforza LL, Oefner PJ (November 2000). "Y chromosome sequence variation and the history of human populations". Nature Genetics. 26 (3): 358–61. doi:10.1038/81685. PMID 11062480.
- ^ Xue Y, Zerjal T, Bao W, Zhu S, Shu Q, Xu J, Du R, Fu S, Li P, Hurles ME, Yang H, Tyler-Smith C (April 2006). "Male demography in East Asia: a north-south contrast in human population expansion times". Genetics. 172 (4): 2431–9. doi:10.1534/genetics.105.054270. PMC 1456369. PMID 16489223.
- ^ Wang CC, Wang LX, Shrestha R, Zhang M, Huang XY, Hu K, Jin L, Li H (2014). "Genetic structure of Qiangic populations residing in the western Sichuan corridor". PLOS ONE. 9 (8): e103772. Bibcode:2014PLoSO...9j3772W. doi:10.1371/journal.pone.0103772. PMC 4121179. PMID 25090432.
{{cite journal}}: CS1 maint: unflagged free DOI (link) - ^ Wen B, Xie X, Gao S, Li H, Shi H, Song X, Qian T, Xiao C, Jin J, Su B, Lu D, Chakraborty R, Jin L (May 2004). "Analyses of genetic structure of Tibeto-Burman populations reveals sex-biased admixture in southern Tibeto-Burmans". American Journal of Human Genetics. 74 (5): 856–65. doi:10.1086/386292. PMC 1181980. PMID 15042512.
- ^ a b Cai X, Qin Z, Wen B, Xu S, Wang Y, Lu Y, Wei L, Wang C, Li S, Huang X, Jin L, Li H (2011). "Human migration through bottlenecks from Southeast Asia into East Asia during Last Glacial Maximum revealed by Y chromosomes". PLOS ONE. 6 (8): e24282. Bibcode:2011PLoSO...624282C. doi:10.1371/journal.pone.0024282. PMC 3164178. PMID 21904623.
{{cite journal}}: CS1 maint: unflagged free DOI (link) - ^ Dongsheng Lu et al. Ancestral Origins and Genetic History of Tibetan Highlanders, August 25, 2016
- ^ Shi, Hong; Zhong, Hua; Peng, Yi; Dong, Yong-Li; Qi, Xue-Bin; Zhang, Feng; Liu, Lu-Fang; Tan, Si-Jie; Ma, Runlin Z (2008-10-29). "Y chromosome evidence of earliest modern human settlement in East Asia and multiple origins of Tibetan and Japanese populations". BMC Biology. 6: 45. doi:10.1186/1741-7007-6-45. ISSN 1741-7007. PMC 2605740. PMID 18959782.
{{cite journal}}: CS1 maint: unflagged free DOI (link) - ^ Kim 2011
- ^ Tajima A, Hayami M, Tokunaga K, Juji T, Matsuo M, Marzuki S, Omoto K, Horai S (2004). "Genetic origins of the Ainu inferred from combined DNA analyses of maternal and paternal lineages". Journal of Human Genetics. 49 (4): 187–93. doi:10.1007/s10038-004-0131-x. PMID 14997363.
- ^ a b YOUICHI SATO, TOSHIKATSU SHINKA, ASHRAF A. EWIS, AIKO YAMAUCHI, TERUAKI IWAMOTO, YUTAKA NAKAHORI Overview of genetic variation in the Y chromosome of modern Japanese males.
- ^ Tumonggor, Meryanne K; Karafet, Tatiana M; Downey, Sean; Lansing, J Stephen; Norquest, Peter; Sudoyo, Herawati; Hammer, Michael F; Cox, Murray P (September 2014). "Isolation, contact and social behavior shaped genetic diversity in West Timor". Journal of Human Genetics. 59 (9): 494–503. doi:10.1038/jhg.2014.62. PMC 4521296. PMID 25078354.
- ^ Cite error: The named reference
Hammerwas invoked but never defined (see the help page). - ^ 崎谷満『DNA・考古・言語の学際研究が示す新・日本列島史』(勉誠出版 2009年)(in Japanese)
- ^ Mondal, Mayukh; Bergström, Anders; Xue, Yali; Calafell, Francesc; Laayouni, Hafid; Casals, Ferran; Majumder, Partha P.; Tyler-Smith, Chris; Bertranpetit, Jaume (25 April 2017). "Y-chromosomal sequences of diverse Indian populations and the ancestry of the Andamanese". Human Genetics. 136 (5): 499–510. doi:10.1007/s00439-017-1800-0. hdl:10230/34399. PMID 28444560.
- ^ Thangaraj K, Singh L, Reddy AG, Rao VR, Sehgal SC, Underhill PA, Pierson M, Frame IG, Hagelberg E (January 2003). "Genetic affinities of the Andaman Islanders, a vanishing human population". Current Biology. 13 (2): 86–93. doi:10.1016/S0960-9822(02)01336-2. PMID 12546781.
- ^ Cordaux R, Weiss G, Saha N, Stoneking M (August 2004). "The northeast Indian passageway: a barrier or corridor for human migrations?". Molecular Biology and Evolution. 21 (8): 1525–33. doi:10.1093/molbev/msh151. PMID 15128876.
- ^ Chandrasekar A, Saheb SY, Gangopadyaya P, Gangopadyaya S, Mukherjee A, Basu D, Lakshmi GR, Sahani AK, Das B, Battacharya S, Kumar S, Xaviour D, Sun D, Rao VR (2007). "YAP insertion signature in South Asia". Annals of Human Biology. 34 (5): 582–6. doi:10.1080/03014460701556262. PMID 17786594.
- ^ Reddy BM, Langstieh BT, Kumar V, Nagaraja T, Reddy AN, Meka A, Reddy AG, Thangaraj K, Singh L (November 2007). "Austro-Asiatic tribes of Northeast India provide hitherto missing genetic link between South and Southeast Asia". PLOS ONE. 2 (11): e1141. Bibcode:2007PLoSO...2.1141R. doi:10.1371/journal.pone.0001141. PMC 2065843. PMID 17989774.
{{cite journal}}: CS1 maint: unflagged free DOI (link) - ^ Y-DNA Haplogroup D and its Subclades - 2019
- ^ Y-DNA Haplogroup D and its Subclades - 2014
- Underhill PA, Kivisild T (2007). "Use of y chromosome and mitochondrial DNA population structure in tracing human migrations". Annual Review of Genetics. 41: 539–64. doi:10.1146/annurev.genet.41.110306.130407. PMID 18076332.
External links
- Atlas of the Human Journey: Genetic Markers, Haplogroup D-M174 (M174), from The Genographic Project at National Geographic
- Famous dna of Japan
