Rifamycin
 | |
 | |
| Clinical data | |
|---|---|
| Trade names | Aemcolo |
| AHFS/Drugs.com | Monograph |
| MedlinePlus | a619010 |
| License data | |
| Routes of administration | By mouth |
| ATC code | |
| Legal status | |
| Legal status |
|
| Identifiers | |
| CAS Number | |
| PubChem CID | |
| DrugBank | |
| ChemSpider | |
| UNII | |
| KEGG | |
| ChEBI | |
| ChEMBL | |
| Chemical and physical data | |
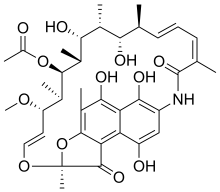
| Formula | C37H47NO12 |
| Molar mass | 697.778 g·mol−1 |
| 3D model (JSmol) | |
| |
| |
The rifamycins are a group of antibiotics that are synthesized either naturally by the bacterium Amycolatopsis rifamycinica or artificially. They are a subclass of the larger family of ansamycins. Rifamycins are particularly effective against mycobacteria, and are therefore used to treat tuberculosis, leprosy, and mycobacterium avium complex (MAC) infections.
The rifamycin group includes the "classic" rifamycin drugs as well as the rifamycin derivatives rifampicin (or rifampin), rifabutin, rifapentine, rifalazil and rifaximin. Rifamycin, sold under the trade name Aemcolo, is approved in the United States for treatment of travelers' diarrhea in some circumstances.[1][2][3]
The name "rifamycin" (originally "rifomycin") was derived from the 1955 French film Rififi.[4]: S402
Bacterium[edit]
Streptomyces mediterranei was first isolated in 1957 from a soil sample collected near the beach-side town of St Raphael in southern France. The name was originally given by two microbiologists working with the Italian drug company Group Lepetit SpA in Milan, the Italian Grazia Beretta, and Pinhas Margalith of Israel.[5]
In 1969, the bacterium was renamed Nocardia mediterranei when another scientist named Thiemann found that it has a cell wall typical of the Nocardia species. Then, in 1986, the bacterium was renamed again Amycolatopsis mediterranei, as the first species of a new genus, because a scientist named Lechevalier discovered that the cell wall lacks mycolic acid and is not able to be infected by the Nocardia and Rhodococcus phages. Based on 16S ribosomal RNA sequences, Bala et al. renamed the species in 2004 Amycolatopsis rifamycinica.
First drugs[edit]
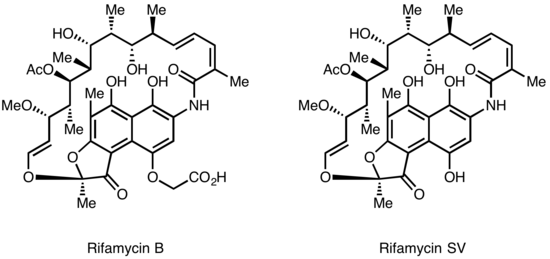
Rifamycins were first isolated in 1957 from a fermentation culture of Streptomyces mediterranei at the laboratory of Gruppo Lepetit SpA in Milan by two scientist named Piero Sensi and Maria Teresa Timbal, working with the Israeli scientist Pinhas Margalith. Initially, a family of closely related antibiotics was discovered referred to as Rifamycin A, B, C, D, E. The only component of this mixture sufficiently stable to isolate in a pure form was Rifamycin B, which unfortunately was poorly active. However, further studies showed that while Rifamycin B was essentially inactive, it was spontaneously oxidized and hydrolyzed in aqueous solutions to yield the highly active Rifamycin S. Simple reduction of Rifamycin S yielded the hydroquinone form called Rifamycin SV, which became the first member of this class to enter clinical use as an intravenous antibiotic. Further chemical modification of Rifamycin SV yielded an improved analog Rifamide, which was also introduced into clinical practice, but was similarly limited to intravenous use. After an extensive modification program, Rifampin was eventually produced, which is orally available and has become a mainstay of Tuberculosis therapy[4]
Lepetit filed for patent protection of Rifamycin B in the UK in August 1958, and in the US in March 1959. The British patent GB921045 was granted in March 1963, and U.S. Patent 3,150,046 was granted in September 1964. The drug is widely regarded as having helped conquer the issue of drug-resistant tuberculosis in the 1960s.
Clinical trials[edit]
Rifamycins have been used for the treatment of many diseases, the most important one being HIV-related tuberculosis. A systematic review of clinical trials on alternative regimens for prevention of active tuberculosis in HIV-negative individuals with latent TB found that a weekly, directly observed regimen of rifapentine with isoniazid for three months was as effective as a daily, self-administered regimen of isoniazid for nine months. But the rifapentine-isoniazid regimen had higher rates of treatment completion and lower rates of hepatotoxicity. However, the rate of treatment-limiting adverse events was higher in the rifapentine-isoniazid regimen.[6]
The rifamycins have a unique mechanism of action, selectively inhibiting bacterial DNA-dependent RNA polymerase, and show no cross-resistance with other antibiotics in clinical use. However, despite their activity against bacteria resistant to other antibiotics, the rifamycins themselves suffer from a rather high frequency of resistance. Because of this, Rifampin and other rifamycins are typically used in combination with other antibacterial drugs. This is routinely practiced in TB therapy and serves to prevent the formation of mutants that are resistant to any of the drugs in the combination. Rifampin rapidly kills fast-dividing bacilli strains as well as "persisters" cells, which remain biologically inactive for long periods of time that allow them to evade antibiotic activity.[7] In addition, rifabutin and rifapentine have both been used against tuberculosis acquired in HIV-positive patients. Although Tuberculosis therapy remains the most important use of Rifampin, an increasing problem with serious Multiple Drug Resistant bacterial infections has led to some use of antibiotic combinations containing Rifampin to treat them.
Mechanism of action[edit]
The antibacterial activity of rifamycins relies on the inhibition of bacterial DNA-dependent RNA synthesis.[8] This is due to the high affinity of rifamycins for the prokaryotic RNA polymerase. The selectivity of the rifamycins depends on the fact that they have a very poor affinity for the analogous mammalian enzyme. Crystal structure data of the antibiotic bound to RNA polymerase indicates that rifamycin blocks synthesis by causing strong steric clashes with the growing oligonucleotide ("steric-occlusion" mechanism).[9][10] If rifamycin binds the polymerase after the chain extension process has started, no inhibition is observed on the biosynthesis, consistent with a steric-occlusion mechanism. Single step high level resistance to the rifamycins occurs as the result of a single amino acid change in the bacterial DNA dependent RNA polymerase.
Biosynthesis[edit]
The first information on the biosynthesis of the rifamycins came from studies using the stable isotope Carbon-13 and NMR spectroscopy to establish the origin of the carbon skeleton. These studies showed that the ansa chain was derived from acetate and propionate, in common with other polyketide antibiotics. The naphthalenic chromophore was shown to derive from a propionate unit coupled with a seven carbon amino moiety of unknown origin. The general scheme of biosynthesis starts with the uncommon starting unit, 3-amino-5-hydroxybenzoic acid (AHBA), via type I polyketide pathway (PKS I) in which chain extension is performed using 2 acetate and 8 propionate units.[11] AHBA is believed to have originated from the Shikimate pathway, however this was not incorporated into the biosynthetic mechanism. This is due to the observation that 3 amino-acid analogues converted into AHBA in cell-free extracts of A. mediterranei.[12]
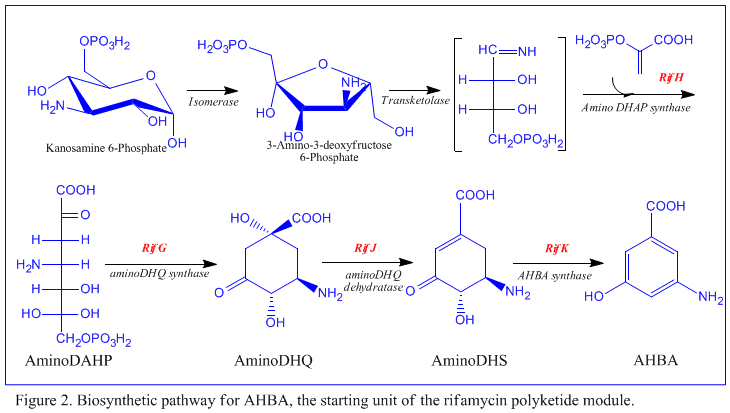
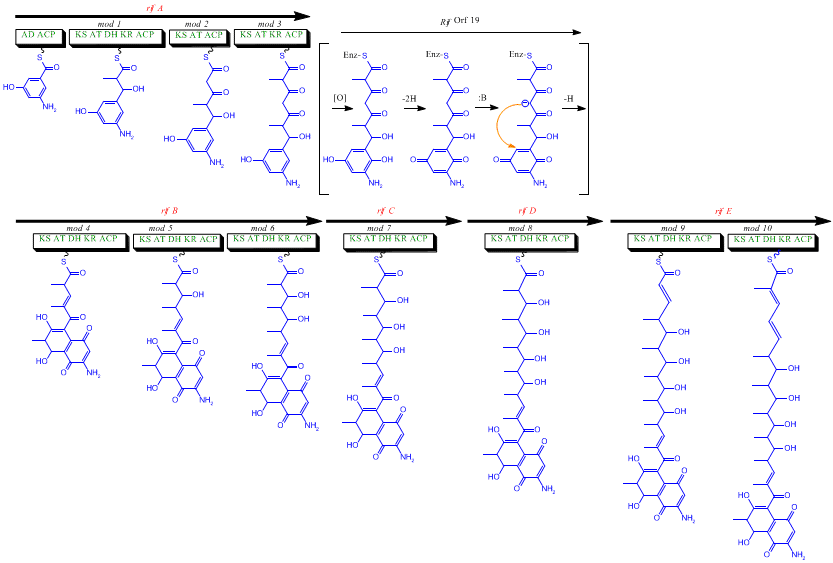

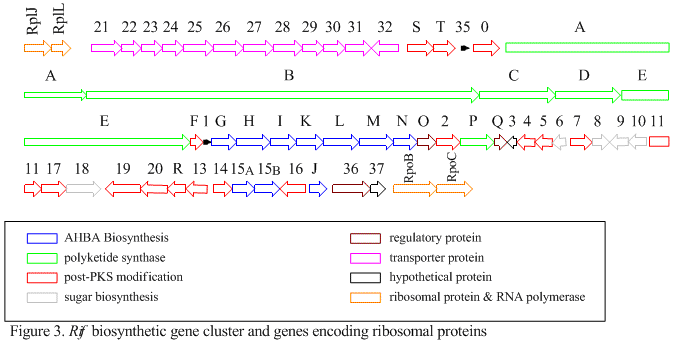
The rif cluster is responsible for the biosynthesis of rifamycins. It contains genes rifG through rifN, which were shown to biosynthesize AHBA.[10] RifK, rifL, rifM, and rifN are believed to act as transaminases in order to form the AHBA precursor kanosamine.[13][14] "RifH" encodes aminoDAHP synthase that catalyzes the condensation between 1-deoxy-1-imino-d-erythrose 4-phosphate and phosphoenolpyruvate.[15] RifA through rifE encode a type I polyketide synthase module, with the loading module being a non-ribosomal peptide synthetase. In all, rifA-E assemble a linear undecaketide and are followed by rifF, which encodes an amide synthase and causes the undecaketide to release and form a macrolactam structure. Moreover, the rif cluster contains various regulatory proteins and glycosylating genes that appear to be silent. Other types of genes seem to perform post-synthase modifications of the original polyketide.
Derivatives[edit]
Lepetit introduced Rifampicin, an orally active rifamycin, in 1966.[16] Rifabutin, a derivative of rifamycin S, was invented by Italian drug manufacturer Achifar in 1975 and came onto the US market in 1992.[16] Hoechst Marion Roussel (now part of Aventis) introduced rifapentine to the US market in 1998, with Achifar having synthesized it in 1965.[17] Use of rifapentine remains uncommon as a treatment for pulmonary tuberculosis, and treatment with rifapentine is given on the basis of careful selection of patients.[18]
Rifaximin is an oral rifamycin marketed in the US by Salix Pharmaceuticals that is poorly absorbed from the intestine. It has been used to treat hepatic encephalopathy and traveler's diarrhea.[19]
Available rifamycins[edit]
- Rifampicin or Rifampin
- Rifabutin
- Rifapentine
- Rifaximin
- Aemcolo [20][21]
References[edit]
- ^ Lin SW, Lin CJ, Yang JC (August 2017). "Rifamycin SV MMX for the treatment of traveler's diarrhea". Expert Opinion on Pharmacotherapy. 18 (12): 1269–1277. doi:10.1080/14656566.2017.1353079. PMID 28697313. S2CID 8853242.
- ^ "FDA approves new drug to treat travelers' diarrhea". U.S. Food and Drug Administration (FDA) (Press release). 16 November 2018. Retrieved 19 November 2018.
- ^ "Drug Approval Package: Aemcolo (rifamycin)". U.S. Food and Drug Administration (FDA). 21 December 2018. Retrieved 27 December 2019.
- ^ a b Sensi, P. (1983). "History of the Development of Rifampin". Clinical Infectious Diseases. 5 (Suppl 3): S402–S406. doi:10.1093/clinids/5.Supplement_3.S402. PMID 6635432.
- ^ Margalith P, Beretta G (1960). "Rifomycin. XI. taxonomic study on streptomyces mediterranei nov. sp". Mycopathologia et Mycologia Applicata. 13 (4): 321–330. doi:10.1007/BF02089930. ISSN 0301-486X. S2CID 23241543.
- ^ Sharma SK, Sharma A, Kadhiravan T, Tharyan P (July 2013). "Rifamycins (rifampicin, rifabutin and rifapentine) compared to isoniazid for preventing tuberculosis in HIV-negative people at risk of active TB". The Cochrane Database of Systematic Reviews. 2013 (7): CD007545. doi:10.1002/14651858.CD007545.pub2. PMC 6532682. PMID 23828580.
- ^ Pozniak AL, Miller R, Ormerod LP (March 1999). "The treatment of tuberculosis in HIV-infected persons". AIDS. 13 (4): 435–445. doi:10.1097/00002030-199907300-00035. PMID 10197371.
- ^ Calvori C, Frontali L, Leoni L, Tecce G (July 1965). "Effect of rifamycin on protein synthesis". Nature. 207 (995): 417–418. Bibcode:1965Natur.207..417C. doi:10.1038/207417a0. PMID 4957347. S2CID 4144738.
- ^ Campbell EA, Korzheva N, Mustaev A, Murakami K, Nair S, Goldfarb A, Darst SA (March 2001). "Structural mechanism for rifampicin inhibition of bacterial rna polymerase". Cell. 104 (6): 901–912. doi:10.1016/S0092-8674(01)00286-0. PMID 11290327. S2CID 8229399.
- ^ Feklistov A, Mekler V, Jiang Q, Westblade LF, Irschik H, Jansen R, et al. (September 2008). "Rifamycins do not function by allosteric modulation of binding of Mg2+ to the RNA polymerase active center". Proceedings of the National Academy of Sciences of the United States of America. 105 (39): 14820–14825. Bibcode:2008PNAS..10514820F. doi:10.1073/pnas.0802822105. PMC 2567451. PMID 18787125.
- ^ Lancini G, Cavalleri B (1997). "Vancomycin and other glycopeptides". In Strohl GR (ed.). Biotechnology of Antibiotics. New York, USA: Marcel Dekker. p. 521.
- ^ Floss HG, Yu TW (February 2005). "Rifamycin-mode of action, resistance, and biosynthesis". Chemical Reviews. 105 (2): 621–632. doi:10.1021/cr030112j. PMID 15700959.
- ^ Guo J, Frost JW (September 2002). "Kanosamine biosynthesis: a likely source of the aminoshikimate pathway's nitrogen atom". Journal of the American Chemical Society. 124 (36): 10642–10643. doi:10.1021/ja026628m. PMID 12207504.
- ^ Arakawa K, Müller R, Mahmud T, Yu TW, Floss HG (September 2002). "Characterization of the early stage aminoshikimate pathway in the formation of 3-amino-5-hydroxybenzoic acid: the RifN protein specifically converts kanosamine into kanosamine 6-phosphate". Journal of the American Chemical Society. 124 (36): 10644–10645. doi:10.1021/ja0206339. PMID 12207505.
- ^ Guo J, Frost JW (January 2002). "Biosynthesis of 1-deoxy-1-imino-D-erythrose 4-phosphate: a defining metabolite in the aminoshikimate pathway". Journal of the American Chemical Society. 124 (4): 528–529. doi:10.1021/ja016963v. PMID 11804477.
- ^ a b Reddy DS, Sinha A, Kumar A, Saini VK (November 2022). "Drug re-engineering and repurposing: A significant and rapid approach to tuberculosis drug discovery". Archiv der Pharmazie. 355 (11): e2200214. doi:10.1002/ardp.202200214. PMID 35841594. S2CID 250582950.
- ^ Guglielmetti L, Günther G, Leu C, Cirillo D, Duarte R, Garcia-Basteiro AL, et al. (May 2022). "Rifapentine access in Europe: growing concerns over key tuberculosis treatment component". The European Respiratory Journal. 59 (5). doi:10.1183/13993003.00388-2022. PMC 9186306. PMID 35589114.
- ^ Munsiff SS, Kambili C, Ahuja SD (December 2006). "Rifapentine for the treatment of pulmonary tuberculosis". Clinical Infectious Diseases. 43 (11): 1468–1475. doi:10.1086/508278. PMID 17083024.
- ^ Ojetti V, Lauritano EC, Barbaro F, Migneco A, Ainora ME, Fontana L, et al. (June 2009). "Rifaximin pharmacology and clinical implications". Expert Opinion on Drug Metabolism & Toxicology. 5 (6): 675–682. doi:10.1517/17425250902973695. PMID 19442033. S2CID 41970585.
- ^ "AEMCOLO (rifamycin) delayed-release tablets, for oral use" (PDF). Aries Pharmaceuticals, Inc. U.S. Food and Drug Administration. 2018.
- ^ "Aemcolo Oral: Uses, Side Effects, Interactions, Pictures, Warnings & Dosing". WebMD.
Further reading[edit]
- Sensi P, Margalith P, Timbal MT (1959). "Rifomycin, a new antibiotic; preliminary report". Il Farmaco; Edizione Scientifica. 14 (2): 146–147. PMID 13639988. - the paper announcing the discovery of the rifamycins.
- Thiemann JE, Zucco G, Pelizza G (1969). "A proposal for the transfer of Streptomyces mediterranei Margalith and Beretta 1960 to the genus Nocardia as Nocardia mediterranea (Margalith and Beretta) comb. nov". Archiv Fur Mikrobiologie. 67 (2): 147–155. doi:10.1007/BF00409680. PMID 5386179. S2CID 19596120. - the paper which renamed Streptomyces mediterranei as Nocardia mediterranei.
- Lechevalier MP, Prauser H, Labeda DP, Ruan JS (January 1986). "Two new genera of nocardioform actinomycetes: Amycolata gen. nov. and Amycolatopsis gen. nov". International Journal of Systematic and Evolutionary Microbiology. 36 (1): 29–37. doi:10.1099/00207713-36-1-29. - the paper which renamed Nocardia mediterranei as Amycolatopsis mediterranei.
- Bala S, Khanna R, Dadhwal M, Prabagaran SR, Shivaji S, Cullum J, Lal R (July 2004). "Reclassification of Amycolatopsis mediterranei DSM 46095 as Amycolatopsis rifamycinica sp. nov". International Journal of Systematic and Evolutionary Microbiology. 54 (Pt 4): 1145–1149. doi:10.1099/ijs.0.02901-0. PMID 15280283. - the paper with the latest name change
