Mimivirus
| Mimivirus | |
|---|---|
| |
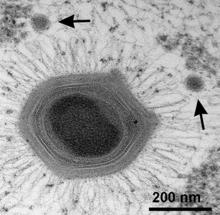
| Mimivirus with two satellite Sputnik virophages (arrows) [1] | |
| Virus classification | |
| (unranked): | Virus |
| Realm: | Varidnaviria |
| Kingdom: | Bamfordvirae |
| Phylum: | Nucleocytoviricota |
| Class: | Megaviricetes |
| Order: | Imitervirales |
| Family: | Mimiviridae |
| Genus: | Mimivirus |
| Species[citation needed] | |

Mimivirus is a genus of giant viruses, in the family Mimiviridae. Amoeba serve as their natural hosts.[2][3] This genus contains a single identified species named Acanthamoeba polyphaga mimivirus (APMV). It also refers to a group of phylogenetically related large viruses.[4]
In colloquial speech, APMV is more commonly referred to as just "mimivirus". Mimivirus, short for "mimicking microbe", is so called to reflect its large size and apparent Gram-staining properties.[5]
Mimivirus has a large and complex genome compared with most other viruses. Until 2013, when a larger virus Pandoravirus was described, it had the largest capsid diameter of all known viruses.[6]
History
[edit]APMV was discovered accidentally in 1992 within the amoeba Acanthamoeba polyphaga, after which it is named, during research into legionellosis by researchers from Marseille and Leeds.[7] The virus was observed in a Gram stain and mistakenly thought to be a Gram-positive bacterium. As a consequence it was named Bradfordcoccus, after Bradford, England, where the amoeba had originated. In 2003, researchers at the Université de la Méditerranée in Marseille, France, published a paper in Science identifying the micro-organism as a virus. It was given the name "mimivirus" (for "mimicking microbe") as it resembles a bacterium on Gram staining.[8]
The same team that discovered the mimivirus later discovered a slightly larger virus, dubbed the mamavirus, and the Sputnik virophage that infects it.[9]
Classification
[edit]Mimivirus has been placed into a viral family by the International Committee on Taxonomy of Viruses as a member of the Mimiviridae,[10] and has been placed into Group I of the Baltimore classification system.[11]
Although not strictly a method of classification, mimivirus joins a group of large viruses known as nucleocytoplasmic large DNA viruses (NCLDV). They are all large viruses which share both molecular characteristics and large genomes. The mimivirus genome also possesses 21 genes encoding homologs to proteins which are seen to be highly conserved in the majority of NCLDVs, and further work suggests that mimivirus is an early divergent of the general NCLDV group.[8]
Structure
[edit]
The mimivirus is the fourth-largest virus, after the Megavirus chilensis, Pandoravirus and Pithovirus. Mimivirus has a capsid diameter of 400 nm. Protein filaments measuring 100 nm project from the surface of the capsid, bringing the total length of the virus up to 600 nm. Variation in scientific literature renders these figures as highly approximate, with the "size" of the virion being casually listed as anywhere between 400 nm and 800 nm, depending on whether total length or capsid diameter is actually quoted.[citation needed]
Its capsid appears hexagonal under an electron microscope, therefore the capsid symmetry is icosahedral.[12] It does not appear to possess an outer viral envelope, suggesting that the virus does not exit the host cell by exocytosis.[13] Mimivirus shares several morphological characteristics with all members of the NCLDV group of viruses. The condensed central core of the virion appears as a dark region under the electron microscope. The large genome of the virus resides within this area. An internal lipid layer surrounding the central core is present in all other NCLDV viruses, so this features may also be present in mimivirus.[12]
Several mRNA transcripts can be recovered from purified virions. Like other NCLDVs, transcripts for DNA polymerase, a capsid protein and a TFII-like transcription factor were found. However, three distinct aminoacyl tRNA synthetase enzyme transcripts and four unknown mRNA molecules specific to mimivirus were also found. These pre-packaged transcripts can be translated without viral gene expression and are likely to be necessary to Mimivirus for replication. Other DNA viruses, such as the Human cytomegalovirus and Herpes simplex virus type-1, also feature pre-packaged mRNA transcripts.[13]
| Genus | Structure | Symmetry | Capsid | Genomic arrangement | Genomic segmentation |
|---|---|---|---|---|---|
| Mimivirus | Icosahedral | T = 972–1141 or T = 1200 (h = 19 ± 1, k = 19 ± 1) | Linear | Monopartite |
Genome
[edit]The mimivirus genome is a linear, double-stranded molecule of DNA with 1,181,404 base pairs in length.[14] This makes it one of the largest viral genomes known, outstripping the next-largest virus genome of the Cafeteria roenbergensis virus by about 450,000 base pairs. In addition, it is larger than at least 30 cellular clades.[15]
In addition to the large size of the genome, mimivirus possesses an estimated 979 protein-coding genes, far exceeding the minimum 4 genes required for viruses to exist (c.f. MS2 and Qβ viruses).[16] Analysis of its genome revealed the presence of genes not seen in any other viruses, including aminoacyl tRNA synthetases, and other genes previously thought only to be encoded by cellular organisms. Like other large DNA viruses, mimivirus contains several genes for sugar, lipid and amino acid metabolism, as well as some metabolic genes not found in any other virus.[13] Roughly 90% of the genome was of coding capacity, with the other 10% being "junk DNA".[citation needed]
Replication
[edit]
A) – C) Surface-shaded rendering of cryoEM reconstruction of untreated Mimivirus
D) The starfish-associated vertex was removed to show the internal nucleocapsid
E) Central slice of the reconstruction looking from the side of the particle
F) Central slice of the reconstruction looking along the 5-fold axis from the starfish-shaped feature
The coloring is based on radial distance from the center of the virus
Gray is from 0 to 1,800 Å
Red from 1,800 to 2,100 Å
Rainbow coloring from red to blue between 2,100 and 2,500 Å
The stages of mimivirus replication are not well known, but as a minimum it is known that mimivirus attaches to a chemical receptor on the surface of an amoeba cell and is taken into the cell. Once inside, an eclipse phase begins, in which the virus disappears and all appears normal within the cell. After about 4 hours small accumulations can be seen in areas of the cell. 8 hours after infection many mimivirus virions are clearly visible within the cell. The cell cytoplasm continues to fill with newly synthesised virions, and about 24 hours after initial infection the cell likely bursts open to release the new mimivirus virions.[13]
Little is known[citation needed][when?] about the details of this replication cycle, most obviously attachment to the cell surface and entry, viral core release, DNA replication, transcription, translation, assembly and release of progeny virions. However, scientists have established the general overview given above using electron micrographs of infected cells.[citation needed] These micrographs show mimivirus capsid assembly in the nucleus, acquisition of an inner lipid membrane via budding from the nucleus, and particles similar to those found in many other viruses, including all NCLDV members. These particles are known in other viruses as viral factories and allow efficient viral assembly by modifying large areas of the host cell.[citation needed]
| Genus | Host details | Tissue tropism | Entry details | Release details | Replication site | Assembly site | Transmission |
|---|---|---|---|---|---|---|---|
| Mimivirus | Zooplankton | None | Unknown | Unknown | Cytoplasm | Nucleus | Passive diffusion |
Pathogenicity
[edit]Mimivirus may be a causative agent of some forms of pneumonia; this is based mainly on indirect evidence in the form of antibodies to the virus discovered in pneumonia patients.[17] However, the classification of mimivirus as a pathogen is tenuous at present as there have been only a couple of papers published potentially linking mimivirus to actual cases of pneumonia. A significant fraction of pneumonia cases are of unknown cause,[18] though a mimivirus has been isolated from a Tunisian woman suffering from pneumonia.[19] There is evidence that mimivirus can infect macrophages.[20]
Implications for defining "life"
[edit]Mimivirus shows many characteristics which place it at the boundary between living and non-living. It is as large as several bacterial species, such as Rickettsia conorii and Tropheryma whipplei, possesses a genomic size comparable to that of several bacteria, including those above, and codes for products previously not thought to be encoded by viruses (including a kind of collagen[21]). In addition, mimivirus has genes coding for nucleotide and amino acid synthesis, which even some small obligate intracellular bacteria lack. They do, however, lack any genes for ribosomal proteins, making mimivirus dependent on a host cell for protein translation and energy metabolism.[citation needed][21]
Because its lineage is very old and could have emerged prior to cellular organisms,[22][23] Mimivirus has added to the debate over the origins of life. Some genes that code for characteristics unique to Mimivirus, including those coding for the capsid, have been conserved in a variety of viruses which infect organisms from all domains. This has been used to suggest that Mimivirus is related to a type of DNA virus that emerged before cellular organisms and played a key role in the development of all life on Earth.[22] An alternative hypothesis is that there were three distinct types of DNA viruses that were involved in generating the three known domains of life—eukarya, archaea and bacteria.[23] It has been suggested that Mimivirus and similar kinds are remnants of a "fourth domain" of life, and that other giant virus may represent other ancient domains.[21]
Nevertheless, mimivirus does not exhibit the following characteristics, all of which are part of many conventional definitions of life:[citation needed]
- homeostasis
- energy metabolism
- response to stimuli
- autopoiesis
- growth via cellular division (instead of replication via self-assembly of individual components)
See also
[edit]- Cafeteria roenbergensis virus a giant marine virus
- Marseillevirus—another giant virus
- Megavirus—another giant virus
- Mycoplasma genitalium—one of the smallest bacteria
- Nanoarchaeum equitans—smallest known independent cell
- Nanobacterium
- Nanobe
- Non-cellular life
- Pandoravirus
- Pithovirus—the largest known virus
- Parvovirus—smallest known viruses
- Pelagibacter ubique—possesses one of the smallest bacterial genomes
- Virophage—a virus that requires the host cell to be co-infected with a giant virus
- The Giant Virus Finder is a software tool that identifies giant viruses in environmental Metagenomes.
References
[edit]- ^ Duponchel, S. and Fischer, M.G. (2019) "Viva lavidaviruses! Five features of virophages that parasitize giant DNA viruses". PLoS pathogens, 15(3). doi:10.1371/journal.ppat.1007592.
 Material was copied from this source, which is available under a Creative Commons Attribution 4.0 International License.
Material was copied from this source, which is available under a Creative Commons Attribution 4.0 International License.
- ^ "Viral Zone". ExPASy. Retrieved 15 June 2015.
- ^ ICTV. "Virus Taxonomy: 2014 Release". Retrieved 15 June 2015.
- ^ Ghedin, E.; Claverie, J. (August 2005). "Mimivirus relatives in the Sargasso sea". Virology Journal. 2: 62. arXiv:q-bio/0504014. Bibcode:2005q.bio.....4014G. doi:10.1186/1743-422X-2-62. PMC 1215527. PMID 16105173.
- ^ Wessner, D. R. (2010). "Discovery of the Giant Mimivirus". Nature Education. 3 (9): 61. Retrieved 7 January 2012.
- ^ "World's biggest virus found in sea off Chile". London: Telegraph UK. 11 October 2011. Archived from the original on 11 October 2011. Retrieved 11 November 2011.
- ^ Richard Birtles; TJ Rowbotham; C Storey; TJ Marrie; Didier Raoult (29 March 1997). "Chlamydia-like obligate parasite of free-living amoebae". The Lancet. 349 (9056): 925–926. doi:10.1016/S0140-6736(05)62701-8. PMID 9093261. S2CID 5382736.
- ^ a b Bernard La Scola; Stéphane Audic; Catherine Robert; Liang Jungang; Xavier de Lamballerie; Michel Drancourt; Richard Birtles; Jean-Michel Claverie; Didier Raoult. (2003). "A giant virus in amoebae". Science. 299 (5615): 2033. doi:10.1126/science.1081867. PMID 12663918. S2CID 39606235.
- ^ Pearson H (2008). "'Virophage' suggests viruses are alive". Nature. 454 (7205): 677. Bibcode:2008Natur.454..677P. doi:10.1038/454677a. ISSN 0028-0836. PMID 18685665.
- ^ Claverie J-M (2010). Mahy W.J. and Van Regenmortel M. H. V. (ed.). Desk Encyclopedia of General Virology (1 ed.). Oxford: Academic Press. p. 189.
- ^ Leppard, Keith; Nigel Dimmock; Easton, Andrew (2007). Introduction to Modern Virology (6 ed.). Blackwell Publishing Limited. pp. 469–470. ISBN 9781405136457.
- ^ a b Xiao C, Kuznetsov YG, Sun S, Hafenstein SL, Kostyuchenko VA, Chipman PR, Suzan-Monti M, Raoult D, McPherson A, Rossmann MG (April 2009). "Structural studies of the giant mimivirus". PLOS Biology. 7 (4): e92. doi:10.1371/journal.pbio.1000092. PMC 2671561. PMID 19402750.
- ^ a b c d Suzan-Monti M, La Scola B, Raoult D (April 2006). "Genomic and evolutionary aspects of Mimivirus". Virus Research. 117 (1): 145–55. doi:10.1016/j.virusres.2005.07.011. PMID 16181700.
- ^ "Acanthamoeba polyphaga mimivirus, complete genome". NCBI.
- ^ Claverie, Jean-Michel; et al. (2006). "Mimivirus and the emerging concept of 'giant' virus". Virus Research. 117 (1): 133–144. arXiv:q-bio/0506007. doi:10.1016/j.virusres.2006.01.008. PMID 16469402. S2CID 8791457.
- ^ Prescott, Lansing M. (1993). Microbiology (2nd ed.). Dubuque, IA: Wm. C. Brown Publishers. ISBN 0-697-01372-3.[page needed]
- ^ La Scola B, Marrie T, Auffray J, Raoult D (2005). "Mimivirus in pneumonia patients". Emerg Infect Dis. 11 (3): 449–52. doi:10.3201/eid1103.040538. PMC 3298252. PMID 15757563. Archived from the original on 24 April 2009. Retrieved 10 September 2017.
- ^ Marrie TJ, Durant H, Yates L (1989). "Community-Acquired Pneumonia Requiring Hospitalization: 5-Year Prospective Study". Reviews of Infectious Diseases. 11 (4): 586–99. doi:10.1093/clinids/11.4.586. PMID 2772465.
- ^ Saadi H, Pagnier I, Colson P, Cherif JK, Beji M, Boughalmi M, Azza S, Armstrong N, Robert C, Fournous G, La Scola B, Raoult D (August 2013). "First isolation of Mimivirus in a patient with pneumonia". Clinical Infectious Diseases. 57 (4): e127–34. doi:10.1093/cid/cit354. PMID 23709652.
- ^ Ghigo, Eric; Kartenbeck, Jürgen; Lien, Pham; Pelkmans, Lucas; Capo, Christian; Mege, Jean-Louis; Raoult, Didier (13 June 2008). "Ameobal Pathogen Mimivirus Infects Macrophages through Phagocytosis". PLOS Pathogens. 4 (6): e1000087. doi:10.1371/journal.ppat.1000087. PMC 2398789. PMID 18551172.
- ^ a b c Garry Hamilton (23 January 2016). "How giant viruses could rewrite the story of life on Earth". New Scientist.
- ^ a b Siebert, Charles (15 March 2006). "Unintelligent Design". Discover Magazine.
- ^ a b Forterre, Patrick (2006). "Three RNA cells for ribosomal lineages and three DNA viruses to replicate their genomes: A hypothesis for the origin of cellular domain". PNAS. 103 (10): 3669–3674. Bibcode:2006PNAS..103.3669F. doi:10.1073/pnas.0510333103. PMC 1450140. PMID 16505372.
Further reading
[edit]- Raoult, D.; et al. (2004). "The 1.2-megabase genome sequence of Mimivirus". Science. 306 (5700): 1344–1350. Bibcode:2004Sci...306.1344R. doi:10.1126/science.1101485. PMID 15486256. S2CID 84298461.
- Ghedin, Elodie; Claverie, J. M. (2005). "Mimivirus relatives in the Sargasso sea". Virology Journal. 2: 62. arXiv:q-bio/0504014. Bibcode:2005q.bio.....4014G. doi:10.1186/1743-422X-2-62. PMC 1215527. PMID 16105173.
- Peplow, Mark, 2004, "Giant virus qualifies as 'living organism'," News@Nature.
- "Mimivirus: discovery of a giant virus". Press Release. Paris: Centre national de la recherche scientifique. 28 March 2003. Archived from the original on 3 June 2004.
- New Scientist, Issue 2544, 25 March 2006.
- Highfield, Roger (15 October 2004). "The Bradford bug that may be a new life form". The Daily Telegraph. London. Archived from the original on 13 March 2007.
- "Scientists investigate structural details of the largest known virus". Science News. 28 April 2009. Archived from the original on 7 January 2012. Retrieved 29 April 2009.
- Keim, Brandon (5 May 2009). "Viral Missing Link Caught on Film". Wired. Wired Science.
External links
[edit]- Viralzone: Mimiviridae
- International Committee on Taxonomy of Viruses (ICTV) picture gallery—images of mimivirus.
- Van Etten, James L. (2011). "Giant Viruses: The recent discovery of really, really big viruses is changing views about the nature of viruses and the history of life". American Scientist. 99 (4): 304. doi:10.1511/2011.91.304. Archived from the original on 10 June 2016. Retrieved 8 July 2011.
- The webpage for Mimivirus
- Radiolab.org Shrink on the discovery of Mimivirus Thursday, July 30, 2015 - 08:54 PM.
