DNA virus
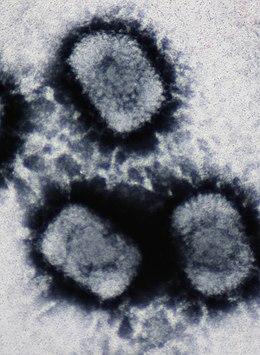
A DNA virus is a virus that has a genome made of deoxyribonucleic acid (DNA) that is replicated by a DNA polymerase. They can be divided between those that have two strands of DNA in their genome, called double-stranded DNA (dsDNA) viruses, and those that have one strand of DNA in their genome, called single-stranded DNA (ssDNA) viruses. dsDNA viruses primarily belong to two realms: Duplodnaviria and Varidnaviria, and ssDNA viruses are almost exclusively assigned to the realm Monodnaviria, which also includes some dsDNA viruses. Additionally, many DNA viruses are unassigned to higher taxa. Reverse transcribing viruses, which have a DNA genome that is replicated through an RNA intermediate by a reverse transcriptase, are classified into the kingdom Pararnavirae in the realm Riboviria.
DNA viruses are ubiquitous worldwide, especially in marine environments where they form an important part of marine ecosystems, and infect both prokaryotes and eukaryotes. They appear to have multiple origins, as viruses in Monodnaviria appear to have emerged from archaeal and bacterial plasmids on multiple occasions, though the origins of Duplodnaviria and Varidnaviria are less clear.
Prominent disease-causing DNA viruses include herpesviruses, papillomaviruses, and poxviruses.
Baltimore classification
[edit]The Baltimore classification system is used to group viruses together based on their manner of messenger RNA (mRNA) synthesis and is often used alongside standard virus taxonomy, which is based on evolutionary history. DNA viruses constitute two Baltimore groups: Group I: double-stranded DNA viruses, and Group II: single-stranded DNA viruses. While Baltimore classification is chiefly based on transcription of mRNA, viruses in each Baltimore group also typically share their manner of replication. Viruses in a Baltimore group do not necessarily share genetic relation or morphology.[1]
Double-stranded DNA viruses
[edit]The first Baltimore group of DNA viruses are those that have a double-stranded DNA genome. All dsDNA viruses have their mRNA synthesized in a three-step process. First, a transcription preinitiation complex binds to the DNA upstream of the site where transcription begins, allowing for the recruitment of a host RNA polymerase. Second, once the RNA polymerase is recruited, it uses the negative strand as a template for synthesizing mRNA strands. Third, the RNA polymerase terminates transcription upon reaching a specific signal, such as a polyadenylation site.[2][3][4]
dsDNA viruses make use of several mechanisms to replicate their genome. Bidirectional replication, in which two replication forks are established at a replication origin site and move in opposite directions of each other, is widely used.[5] A rolling circle mechanism that produces linear strands while progressing in a loop around the circular genome is also common.[6][7] Some dsDNA viruses use a strand displacement method whereby one strand is synthesized from a template strand, and a complementary strand is then synthesized from the prior synthesized strand, forming a dsDNA genome.[8] Lastly, some dsDNA viruses are replicated as part of a process called replicative transposition whereby a viral genome in a host cell's DNA is replicated to another part of a host genome.[9]
dsDNA viruses can be subdivided between those that replicate in the cell nucleus, and as such are relatively dependent on host cell machinery for transcription and replication, and those that replicate in the cytoplasm, in which case they have evolved or acquired their own means of executing transcription and replication.[10] dsDNA viruses are also commonly divided between tailed dsDNA viruses, referring to members of the realm Duplodnaviria, usually the tailed bacteriophages of the order Caudovirales, and tailless or non-tailed dsDNA viruses of the realm Varidnaviria.[11][12]
Single-stranded DNA viruses
[edit]
The second Baltimore group of DNA viruses are those that have a single-stranded DNA genome. ssDNA viruses have the same manner of transcription as dsDNA viruses. However, because the genome is single-stranded, it is first made into a double-stranded form by a DNA polymerase upon entering a host cell. mRNA is then synthesized from the double-stranded form. The double-stranded form of ssDNA viruses may be produced either directly after entry into a cell or as a consequence of replication of the viral genome.[13][14] Eukaryotic ssDNA viruses are replicated in the nucleus.[10][15]
Most ssDNA viruses contain circular genomes that are replicated via rolling circle replication (RCR). ssDNA RCR is initiated by an endonuclease that bonds to and cleaves the positive strand, allowing a DNA polymerase to use the negative strand as a template for replication. Replication progresses in a loop around the genome by means of extending the 3'-end of the positive strand, displacing the prior positive strand, and the endonuclease cleaves the positive strand again to create a standalone genome that is ligated into a circular loop. The new ssDNA may be packaged into virions or replicated by a DNA polymerase to form a double-stranded form for transcription or continuation of the replication cycle.[13][16]
Parvoviruses contain linear ssDNA genomes that are replicated via rolling hairpin replication (RHR), which is similar to RCR. Parvovirus genomes have hairpin loops at each end of the genome that repeatedly unfold and refold during replication to change the direction of DNA synthesis to move back and forth along the genome, producing numerous copies of the genome in a continuous process. Individual genomes are then excised from this molecule by the viral endonuclease. For parvoviruses, either the positive or negative sense strand may be packaged into capsids, varying from virus to virus.[16][17]
Nearly all ssDNA viruses have positive sense genomes, but a few exceptions and peculiarities exist. The family Anelloviridae is the only ssDNA family whose members have negative sense genomes, which are circular.[15] Parvoviruses, as previously mentioned, may package either the positive or negative sense strand into virions.[14] Lastly, bidnaviruses package both the positive and negative linear strands.[15][18]
ICTV classification
[edit]The International Committee on Taxonomy of Viruses (ICTV) oversees virus taxonomy and organizes viruses at the basal level at the rank of realm. Virus realms correspond to the rank of domain used for cellular life but differ in that viruses within a realm do not necessarily share common ancestry, nor do the realms share common ancestry with each other. As such, each virus realm represents at least one instance of viruses coming into existence. Within each realm, viruses are grouped together based on shared characteristics that are highly conserved over time.[19] Three DNA virus realms are recognized: Duplodnaviria, Monodnaviria, and Varidnaviria.
Duplodnaviria
[edit]
Duplodnaviria contains dsDNA viruses that encode a major capsid protein (MCP) that has the HK97 fold. Viruses in the realm also share a number of other characteristics involving the capsid and capsid assembly, including an icosahedral capsid shape and a terminase enzyme that packages viral DNA into the capsid during assembly. Two groups of viruses are included in the realm: tailed bacteriophages, which infect prokaryotes and are assigned to the order Caudovirales, and herpesviruses, which infect animals and are assigned to the order Herpesvirales.[11]
Duplodnaviria is a very ancient realm, perhaps predating the last universal common ancestor (LUCA) of cellular life. Its origins not known, nor whether it is monophyletic or polyphyletic. A characteristic feature is the HK97-fold found in the MCP of all members, which is found outside the realm only in encapsulins, a type of nanocompartment found in bacteria: this relation is not fully understood.[11][20][21]
The relation between caudoviruses and herpesviruses is also uncertain: they may share a common ancestor or herpesviruses may be a divergent clade from the realm Caudovirales. A common trait among duplodnaviruses is that they cause latent infections without replication while still being able to replicate in the future.[22][23] Tailed bacteriophages are ubiquitous worldwide,[24] important in marine ecology,[25] and the subject of much research.[26] Herpesviruses are known to cause a variety of epithelial diseases, including herpes simplex, chickenpox and shingles, and Kaposi's sarcoma.[27][28][29]
Monodnaviria
[edit]Monodnaviria contains ssDNA viruses that encode an endonuclease of the HUH superfamily that initiates rolling circle replication and all other viruses descended from such viruses. The prototypical members of the realm are called CRESS-DNA viruses and have circular ssDNA genomes. ssDNA viruses with linear genomes are descended from them, and in turn some dsDNA viruses with circular genomes are descended from linear ssDNA viruses.[30]
Viruses in Monodnaviria appear to have emerged on multiple occasions from archaeal and bacterial plasmids, a type of extra-chromosomal DNA molecule that self-replicates inside its host. The kingdom Shotokuvirae in the realm likely emerged from recombination events that merged the DNA of these plasmids and complementary DNA encoding the capsid proteins of RNA viruses.[30][31]
CRESS-DNA viruses include three kingdoms that infect prokaryotes: Loebvirae, Sangervirae, and Trapavirae. The kingdom Shotokuvirae contains eukaryotic CRESS-DNA viruses and the atypical members of Monodnaviria.[30] Eukaryotic monodnaviruses are associated with many diseases, and they include papillomaviruses and polyomaviruses, which cause many cancers,[32][33] and geminiviruses, which infect many economically important crops.[34]
Varidnaviria
[edit]
Varidnaviria contains DNA viruses that encode MCPs that have a jelly roll fold folded structure in which the jelly roll (JR) fold is perpendicular to the surface of the viral capsid. Many members also share a variety of other characteristics, including a minor capsid protein that has a single JR fold, an ATPase that packages the genome during capsid assembly, and a common DNA polymerase. Two kingdoms are recognized: Helvetiavirae, whose members have MCPs with a single vertical JR fold, and Bamfordvirae, whose members have MCPs with two vertical JR folds.[12]
Varidnaviria is either monophyletic or polyphyletic and may predate the LUCA. The kingdom Bamfordvirae is likely derived from the other kingdom Helvetiavirae via fusion of two MCPs to have an MCP with two jelly roll folds instead of one. The single jelly roll (SJR) fold MCPs of Helvetiavirae show a relation to a group of proteins that contain SJR folds, including the Cupin superfamily and nucleoplasmins.[12][20][21]
Marine viruses in Varidnaviria are ubiquitous worldwide and, like tailed bacteriophages, play an important role in marine ecology.[35] Most identified eukaryotic DNA viruses belong to the realm.[36] Notable disease-causing viruses in Varidnaviria include adenoviruses, poxviruses, and the African swine fever virus.[37] Poxviruses have been highly prominent in the history of modern medicine, especially Variola virus, which caused smallpox.[38] Many varidnaviruses can become endogenized in their host's genome; a peculiar example are virophages, which after infecting a host, can protect the host against giant viruses.[36]
Baltimore classification
[edit]dsDNA viruses are classified into three realms and include many taxa that are unassigned to a realm:
- All viruses in Duplodnaviria are dsDNA viruses.[11]
- In Monodnaviria, members of the class Papovaviricetes are dsDNA viruses.[30]
- All viruses in Varidnaviria are dsDNA viruses.[12]
- The following taxa that are unassigned to a realm exclusively contain dsDNA viruses:[12]
- Orders: Ligamenvirales
- Families: Ampullaviridae, Baculoviridae, Bicaudaviridae, Clavaviridae, Fuselloviridae, Globuloviridae, Guttaviridae, Halspiviridae, Hytrosaviridae, Nimaviridae, Nudiviridae, Ovaliviridae, Plasmaviridae, Polydnaviridae, Portogloboviridae, Thaspiviridae, Tristromaviridae
- Genera: Dinodnavirus, Rhizidiovirus
ssDNA viruses are classified into one realm and include several families that are unassigned to a realm:
- In Monodnaviria, all members except viruses in Papovaviricetes are ssDNA viruses.[30]
- The unassigned families Anelloviridae and Spiraviridae are ssDNA virus families.[30]
- Viruses in the family Finnlakeviridae contain ssDNA genomes. Finnlakeviridae is unassigned to a realm but is a proposed member of Varidnaviria.[12]
References
[edit]- ^ Lostroh 2019, pp. 11–13
- ^ "dsDNA templated transcription". ViralZone. Swiss Institute of Bioinformatics. Retrieved 24 September 2020.
- ^ Rampersad 2018, p. 66
- ^ Fermin 2018, pp. 36–40
- ^ "dsDNA bidirectional replication". ViralZone. Swiss Institute of Bioinformatics. Retrieved 24 September 2020.
- ^ "dsDNA rolling circle replication". ViralZone. Swiss Institute of Bioinformatics. Retrieved 24 September 2020.
- ^ Bernstein H, Bernstein C (5 July 1973). "Circular and branched circular concatenates as possible intermediates in bacteriophage T4 DNA replication". J Mol Biol. 77 (3): 355–361. doi:10.1016/0022-2836(73)90443-9. PMID 4580243.
- ^ "DNA strand displacement replication". ViralZone. Swiss Institute of Bioinformatics. Retrieved 24 September 2020.
- ^ "Replicative transposition". ViralZone. Swiss Institute of Bioinformatics. Retrieved 24 September 2020.
- ^ a b Cann 2015, pp. 122–127
- ^ a b c d Koonin EV, Dolja VV, Krupovic M, Varsani A, Wolf YI, Yutin N, Zerbini M, Kuhn JH (18 October 2019). "Create a megataxonomic framework, filling all principal/primary taxonomic ranks, for dsDNA viruses encoding HK97-type major capsid proteins" (docx). International Committee on Taxonomy of Viruses. Retrieved 24 September 2020.
- ^ a b c d e f Koonin EV, Dolja VV, Krupovic M, Varsani A, Wolf YI, Yutin N, Zerbini M, Kuhn JH (18 October 2019). "Create a megataxonomic framework, filling all principal taxonomic ranks, for DNA viruses encoding vertical jelly roll-type major capsid proteins" (docx). International Committee on Taxonomy of Viruses. Retrieved 24 September 2020.
- ^ a b "ssDNA Rolling circle". ViralZone. Swiss Institute of Bioinformatics. Retrieved 24 September 2020.
- ^ a b "Rolling hairpin replication". ViralZone. Swiss Institute of Bioinformatics. Retrieved 24 September 2020.
- ^ a b c Fermin 2018, pp. 40–41
- ^ a b Rampersad 2018, pp. 61–62
- ^ Kerr J, Cotmore S, Bloom ME (25 November 2005). Parvoviruses. CRC Press. pp. 171–185. ISBN 9781444114782.
- ^ "Bidnaviridae". ViralZone. Swiss Institute of Bioinformatics. Retrieved 24 September 2020.
- ^ International Committee on Taxonomy of Viruses Executive Committee (May 2020). "The New Scope of Virus Taxonomy: Partitioning the Virosphere Into 15 Hierarchical Ranks". Nat Microbiol. 5 (5): 668–674. doi:10.1038/s41564-020-0709-x. PMC 7186216. PMID 32341570.
- ^ a b Krupovic M, Koonin EV (21 March 2017). "Multiple origins of viral capsid proteins from cellular ancestors". Proc Natl Acad Sci U S A. 114 (12): E2401 – E2410. Bibcode:2017PNAS..114E2401K. doi:10.1073/pnas.1621061114. PMC 5373398. PMID 28265094.
- ^ a b Krupovic, M; Dolja, VV; Koonin, EV (14 July 2020). "The LUCA and its complex virome" (PDF). Nat Rev Microbiol. 18 (11): 661–670. doi:10.1038/s41579-020-0408-x. PMID 32665595. S2CID 220516514. Archived (PDF) from the original on 27 October 2020. Retrieved 24 September 2020.
- ^ Weidner-Glunde M, Kruminis-Kaszkiel E, Savanagoudar M (February 2020). "Herpesviral Latency—Common Themes". Pathogens. 9 (2): 125. doi:10.3390/pathogens9020125. PMC 7167855. PMID 32075270.
- ^ "Virus latency". ViralZone. Swiss Institute of Bioinformatics. Retrieved 24 September 2020.
- ^ Andrade-Martínez JS, Moreno-Gallego JL, Reyes A (August 2019). "Defining a Core Genome for the Herpesvirales and Exploring their Evolutionary Relationship with the Caudovirales". Sci Rep. 9 (1): 11342. Bibcode:2019NatSR...911342A. doi:10.1038/s41598-019-47742-z. PMC 6683198. PMID 31383901.
- ^ Wilhelm SW, Suttle CA (October 1999). "Viruses and Nutrient Cycles in the Sea: Viruses play critical roles in the structure and function of aquatic food webs". BioScience. 49 (10): 781–788. doi:10.2307/1313569. JSTOR 1313569.
- ^ Keen EC (January 2015). "A century of phage research: Bacteriophages and the shaping of modern biology". BioEssays. 37 (1): 6–9. doi:10.1002/bies.201400152. PMC 4418462. PMID 25521633.
- ^ Kukhanova MK, Korovina AN, Kochetkov SN (December 2014). "Human herpes simplex virus: life cycle and development of inhibitors". Biochemistry (Mosc). 79 (13): 1635–1652. doi:10.1134/S0006297914130124. PMID 25749169. S2CID 7414402.
- ^ Gershon AA, Breuer J, Cohen JI, Cohrs RJ, Gershon MD, Gilden D, Grose C, Hambleton S, Kennedy PG, Oxman MN, Seward JF, Yamanishi K (2 July 2015). "Varicella zoster virus infection". Nat Rev Dis Primers. 1: 15016. doi:10.1038/nrdp.2015.16. PMC 5381807. PMID 27188665.
- ^ O'Leary JJ, Kennedy MM, McGee JO (February 1997). "Kaposi's sarcoma associated herpes virus (KSHV/HHV 8): epidemiology, molecular biology and tissue distribution". Mol Pathol. 50 (1): 4–8. doi:10.1136/mp.50.1.4. PMC 379571. PMID 9208806.
- ^ a b c d e f Koonin EV, Dolja VV, Krupovic M, Varsani A, Wolf YI, Yutin N, Zerbini M, Kuhn JH (18 October 2019). "Create a megataxonomic framework, filling all principal taxonomic ranks, for ssDNA viruses" (docx). International Committee on Taxonomy of Viruses. Retrieved 24 September 2020.
- ^ Kazlauskas D, Varsani A, Koonin EV, Krupovic M (31 July 2019). "Multiple Origins of Prokaryotic and Eukaryotic Single-Stranded DNA Viruses From Bacterial and Archaeal Plasmids". Nat Commun. 10 (1): 3425. Bibcode:2019NatCo..10.3425K. doi:10.1038/s41467-019-11433-0. PMC 6668415. PMID 31366885.
- ^ "Papillomaviridae". ViralZone. Swiss Institute of Bioinformatics. Retrieved 24 September 2020.
- ^ "Polyomaviridae". ViralZone. Swiss Institute of Bioinformatics. Retrieved 24 September 2020.
- ^ Malathi VG, Renuka Devi P (March 2019). "ssDNA Viruses: Key Players in Global Virome". VirusDisease. 30 (1): 3–12. doi:10.1007/s13337-019-00519-4. PMC 6517461. PMID 31143827.
- ^ Kauffman KM, Hussain FA, Yang J, Arevalo P, Brown JM, Chang WK, VanInsberghe D, Elsherbini J, Sharma RS, Cutler MB, Kelly L, Polz MF (1 February 2018). "A Major Lineage of Non-Tailed dsDNA Viruses as Unrecognized Killers of Marine Bacteria". Nature. 554 (7690): 118–122. Bibcode:2018Natur.554..118K. doi:10.1038/nature25474. PMID 29364876. S2CID 4462007.
- ^ a b Krupovic M, Koonin EV (February 2015). "Polintons: a hotbed of eukaryotic virus, transposon and plasmid evolution". Nat Rev Microbiol. 13 (2): 105–115. doi:10.1038/nrmicro3389. PMC 5898198. PMID 25534808.
- ^ "Virus Taxonomy: 2019 Release". International Committee on Taxonomy of Viruses. Retrieved 24 September 2020.
- ^ Meyer H, Ehmann R, Smith GL (February 2020). "Smallpox in the Post-Eradication Era". Viruses. 12 (2): 138. doi:10.3390/v12020138. PMC 7077202. PMID 31991671.
Bibliography
[edit]- Lostroh, P. (2019). Molecular and Cellular Biology of Viruses. Garland Science. ISBN 978-0429664304. Retrieved 24 September 2020.
- Cann, A. (2015). Principles of Molecular Virology. Elsevier. pp. 122–127. ISBN 978-0128019559.
- Fermin, G. (2018). "Virion Structure, Genome Organization, and Taxonomy of Viruses". In Tennant, P.; Fermin, G.; Foster, J. (eds.). Viruses: Molecular Biology, Host Interactions and Applications to Biotechnology. San Diego, CA: Elsevier. pp. 35–46. doi:10.1016/B978-0-12-811257-1.00002-4. ISBN 978-0128112571. S2CID 89706800. Retrieved 8 December 2020.
- Rampersad, S.; Tennant, P. (2018). "Replication and Expression Strategies of Viruses". In Tennant, P.; Fermin, G.; Foster, J. (eds.). Viruses: Molecular Biology, Host Interactions, and Applications to Biotechnology. San Diego, CA: Elsevier. pp. 55–82. doi:10.1016/B978-0-12-811257-1.00003-6. ISBN 978-0128112571. S2CID 90170103. Retrieved 8 December 2020.
