Histidine
 Structure of Histidine
| |||
| |||
| Names | |||
|---|---|---|---|
| IUPAC name
Histidine
| |||
| Other names
2-Amino-3-(1H-imidazol-4-yl)propanoic acid
| |||
| Identifiers | |||
3D model (JSmol)
|
| ||
| 84088 | |||
| ChEBI | |||
| ChEMBL | |||
| ChemSpider | |||
| DrugBank | |||
| ECHA InfoCard | 100.000.678 | ||
| EC Number |
| ||
| 83042 | |||
| KEGG | |||
PubChem CID
|
|||
| UNII | |||
CompTox Dashboard (EPA)
|
|||
| |||
| |||
| Properties | |||
| C6H9N3O2 | |||
| Molar mass | 155.157 g·mol−1 | ||
| 4.19g/100g @ 25 °C [1] | |||
| Hazards | |||
| NFPA 704 (fire diamond) | |||
| Supplementary data page | |||
| Histidine (data page) | |||
Except where otherwise noted, data are given for materials in their standard state (at 25 °C [77 °F], 100 kPa).
| |||
Histidine (symbol His or H)[2] is an essential amino acid that is used in the biosynthesis of proteins. It contains an α-amino group (which is in the protonated –NH3+ form under biological conditions), a carboxylic acid group (which is in the deprotonated –COO− form under biological conditions), and an imidazole side chain (which is partially protonated), classifying it as a positively charged amino acid at physiological pH. Initially thought essential only for infants, it has now been shown in longer-term studies to be essential for adults also.[3] It is encoded by the codons CAU and CAC.
Histidine was first isolated by Albrecht Kossel and Sven Gustaf Hedin in 1896.[4] The name stems from its discovery in tissue, from ἱστός histós "tissue".[2] It is also a precursor to histamine, a vital inflammatory agent in immune responses. The acyl radical is histidyl.
Properties of the imidazole side chain
[edit]The conjugate acid (protonated form) of the imidazole side chain in histidine has a pKa of approximately 6.0. Thus, below a pH of 6, the imidazole ring is mostly protonated (as described by the Henderson–Hasselbalch equation). The resulting imidazolium ring bears two NH bonds and has a positive charge. The positive charge is equally distributed between both nitrogens and can be represented with two equally important resonance structures. Sometimes, the symbol Hip is used for this protonated form instead of the usual His.[5][6][7] Above pH 6, one of the two protons is lost. The remaining proton of the imidazole ring can reside on either nitrogen, giving rise to what are known as the N3-H or N1-H tautomers. The N3-H tautomer is shown in the figure above. In the N1-H tautomer, the NH is nearer the backbone. These neutral tautomers, also referred to as Nε and Nδ, are sometimes referred to with symbols Hie and Hid, respectively.[5][6][7] The imidazole/imidazolium ring of histidine is aromatic at all pH values.[8] Under certain conditions, all three ion-forming groups of histidine can be charged forming the histidinium cation.[9]
The acid-base properties of the imidazole side chain are relevant to the catalytic mechanism of many enzymes.[10] In catalytic triads, the basic nitrogen of histidine abstracts a proton from serine, threonine, or cysteine to activate it as a nucleophile. In a histidine proton shuttle, histidine is used to quickly shuttle protons. It can do this by abstracting a proton with its basic nitrogen to make a positively charged intermediate and then use another molecule, a buffer, to extract the proton from its acidic nitrogen. In carbonic anhydrases, a histidine proton shuttle is utilized to rapidly shuttle protons away from a zinc-bound water molecule to quickly regenerate the active form of the enzyme. In helices E and F of hemoglobin, histidine influences binding of dioxygen as well as carbon monoxide. This interaction enhances the affinity of Fe(II) for O2 but destabilizes the binding of CO, which binds only 200 times stronger in hemoglobin, compared to 20,000 times stronger in free heme.
The tautomerism and acid-base properties of the imidazole side chain has been characterized by 15N NMR spectroscopy. The two 15N chemical shifts are similar (about 200 ppm, relative to nitric acid on the sigma scale, on which increased shielding corresponds to increased chemical shift). NMR spectral measurements shows that the chemical shift of N1-H drops slightly, whereas the chemical shift of N3-H drops considerably (about 190 vs. 145 ppm). This change indicates that the N1-H tautomer is preferred, possibly due to hydrogen bonding to the neighboring ammonium. The shielding at N3 is substantially reduced due to the second-order paramagnetic effect, which involves a symmetry-allowed interaction between the nitrogen lone pair and the excited π* states of the aromatic ring. At pH > 9, the chemical shifts of N1 and N3 are approximately 185 and 170 ppm.[11]
Ligand
[edit]

Histidine forms complexes with many metal ions. The imidazole sidechain of the histidine residue commonly serves as a ligand in metalloproteins. One example is the axial base attached to Fe in myoglobin and hemoglobin. Poly-histidine tags (of six or more consecutive H residues) are utilized for protein purification by binding to columns with nickel or cobalt, with micromolar affinity.[12] Natural poly-histidine peptides, found in the venom of the viper Atheris squamigera have been shown to bind Zn(2+), Ni(2+) and Cu(2+) and affect the function of venom metalloproteases.[13]
Metabolism
[edit]Biosynthesis
[edit]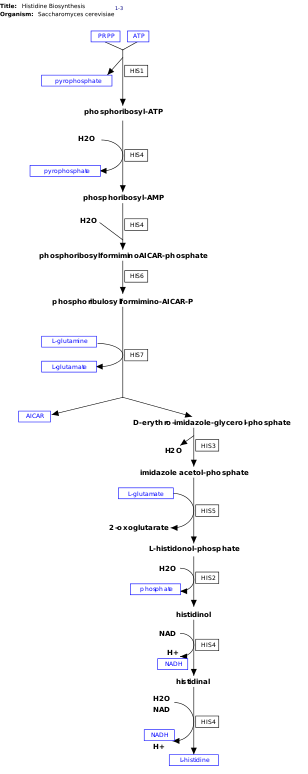
l-Histidine is an essential amino acid that is not synthesized de novo in humans.[14] Humans and other animals must ingest histidine or histidine-containing proteins. The biosynthesis of histidine has been widely studied in prokaryotes such as E. coli. Histidine synthesis in E. coli involves eight gene products (His1, 2, 3, 4, 5, 6, 7, and 8) and it occurs in ten steps. This is possible because a single gene product has the ability to catalyze more than one reaction. For example, as shown in the pathway, His4 catalyzes 4 different steps in the pathway.[15]
Histidine is synthesized from phosphoribosyl pyrophosphate (PRPP), which is made from ribose-5-phosphate by ribose-phosphate diphosphokinase in the pentose phosphate pathway. The first reaction of histidine biosynthesis is the condensation of PRPP and adenosine triphosphate (ATP) by the enzyme ATP-phosphoribosyl transferase. ATP-phosphoribosyl transferase is indicated by His1 in the image.[15] His4 gene product then hydrolyzes the product of the condensation, phosphoribosyl-ATP, producing phosphoribosyl-AMP (PRAMP), which is an irreversible step. His4 then catalyzes the formation of phosphoribosylformiminoAICAR-phosphate, which is then converted to phosphoribulosylformimino-AICAR-P by the His6 gene product.[16] His7 splits phosphoribulosylformimino-AICAR-P to form d-erythro-imidazole-glycerol-phosphate. After, His3 forms imidazole acetol-phosphate releasing water. His5 then makes l-histidinol-phosphate, which is then hydrolyzed by His2 making histidinol. His4 catalyzes the oxidation of l-histidinol to form l-histidinal, an amino aldehyde. In the last step, l-histidinal is converted to l-histidine.[16][17]
The histidine biosynthesis pathway has been studied in the fungus Neurospora crassa, and a gene (His-3) encoding a multienzyme complex was found that was similar to the His4 gene of the bacterium E. coli.[18] A genetic study of N. crassa histidine mutants indicated that the individual activities of the multienzyme complex occur in discrete, contiguous sections of the His-3 genetic map, suggesting that the different activities of the multienzyme complex are encoded separately from each other.[18] However, mutants were also found that lacked all three activities simultaneously, suggesting that some mutations cause loss of function of the complex as a whole.
Just like animals and microorganisms, plants need histidine for their growth and development.[10] Microorganisms and plants are similar in that they can synthesize histidine.[19] Both synthesize histidine from the biochemical intermediate phosphoribosyl pyrophosphate. In general, the histidine biosynthesis is very similar in plants and microorganisms.[20]
Regulation of biosynthesis
[edit]This pathway requires energy in order to occur therefore, the presence of ATP activates the first enzyme of the pathway, ATP-phosphoribosyl transferase (shown as His1 in the image on the right). ATP-phosphoribosyl transferase is the rate determining enzyme, which is regulated through feedback inhibition meaning that it is inhibited in the presence of the product, histidine.[21]
Degradation
[edit]Histidine is one of the amino acids that can be converted to intermediates of the tricarboxylic acid (TCA) cycle (also known as the citric acid cycle).[22] Histidine, along with other amino acids such as proline and arginine, takes part in deamination, a process in which its amino group is removed. In prokaryotes, histidine is first converted to urocanate by histidase. Then, urocanase converts urocanate to 4-imidazolone-5-propionate. Imidazolonepropionase catalyzes the reaction to form formiminoglutamate (FIGLU) from 4-imidazolone-5-propionate.[23] The formimino group is transferred to tetrahydrofolate, and the remaining five carbons form glutamate.[22] Overall, these reactions result in the formation of glutamate and ammonia.[24] Glutamate can then be deaminated by glutamate dehydrogenase or transaminated to form α-ketoglutarate.[22]
Conversion to other biologically active amines
[edit]- The histidine amino acid is a precursor for histamine, an amine produced in the body necessary for inflammation.[25]
- The enzyme histidine ammonia-lyase converts histidine into ammonia and urocanic acid. A deficiency in this enzyme is present in the rare metabolic disorder histidinemia, producing urocanic aciduria as a key diagnostic finding.
- Histidine can be converted to 3-methylhistidine, which serves as a biomarker for skeletal muscle damage, by certain methyltransferase enzymes.[26]
- Histidine is also a precursor for carnosine biosynthesis, which is a dipeptide found in skeletal muscle.[27]
- In Actinomycetota and filamentous fungi, such as Neurospora crassa, histidine can be converted into the antioxidant ergothioneine.[28]

Requirements
[edit]The Food and Nutrition Board (FNB) of the U.S. Institute of Medicine set Recommended Dietary Allowances (RDAs) for essential amino acids in 2002. For histidine, for adults 19 years and older, 14 mg/kg body weight/day.[29] Supplemental histidine is being investigated for use in a variety of different conditions, including neurological disorders, atopic dermatitis, metabolic syndrome, diabetes, uraemic anaemia, ulcers, inflammatory bowel diseases, malignancies, and muscle performance during strenuous exercise.[30]
See also
[edit]References
[edit]- ^ http://prowl.rockefeller.edu/aainfo/solub.htm[full citation needed]
- ^ a b "Nomenclature and Symbolism for Amino Acids and Peptides". IUPAC-IUB Joint Commission on Biochemical Nomenclature. 1983. Archived from the original on 9 October 2008. Retrieved 5 March 2018.
- ^ Kopple, J D; Swendseid, M E (1975). "Evidence that histidine is an essential amino acid in normal and chronically uremic man". Journal of Clinical Investigation. 55 (5): 881–91. doi:10.1172/JCI108016. PMC 301830. PMID 1123426.
- ^ Vickery, Hubert Bradford; Leavenworth, Charles S. (1928-08-01). "On the Separation of Histidine and Arginine" (PDF). Journal of Biological Chemistry. 78 (3): 627–635. doi:10.1016/S0021-9258(18)83967-9. ISSN 0021-9258.
- ^ a b Kim, Meekyum Olivia; Nichols, Sara E.; Wang, Yi; McCammon, J. Andrew (March 2013). "Effects of histidine protonation and rotameric states on virtual screening of M. tuberculosis RmlC". Journal of Computer-Aided Molecular Design. 27 (3): 235–246. Bibcode:2013JCAMD..27..235K. doi:10.1007/s10822-013-9643-9. ISSN 0920-654X. PMC 3639364. PMID 23579613.
- ^ a b "HISTIDINE". ambermd.org. Retrieved 2022-05-12.
- ^ a b Dokainish, Hisham M.; Kitao, Akio (2016-08-05). "Computational Assignment of the Histidine Protonation State in (6-4) Photolyase Enzyme and Its Effect on the Protonation Step". ACS Catalysis. 6 (8): 5500–5507. doi:10.1021/acscatal.6b01094. ISSN 2155-5435. S2CID 88813605.
- ^ Mrozek, Agnieszka; Karolak-Wojciechowska, Janina; Kieć-Kononowicz, Katarzyna (2003). "Five-membered heterocycles. Part III. Aromaticity of 1,3-imidazole in 5+n hetero-bicyclic molecules". Journal of Molecular Structure. 655 (3): 397–403. Bibcode:2003JMoSt.655..397M. doi:10.1016/S0022-2860(03)00282-5.
- ^ Novikov, Anton P.; Safonov, Alexey V.; German, Konstantin E.; Grigoriev, Mikhail S. (2023-12-01). "What kind of interactions we may get moving from zwitter to "dritter" ions: C–O⋯Re(O4) and Re–O⋯Re(O4) anion⋯anion interactions make structural difference between L-histidinium perrhenate and pertechnetate". CrystEngComm. 26: 61–69. doi:10.1039/D3CE01164J. ISSN 1466-8033. S2CID 265572280.
- ^ a b Ingle, Robert A. (2011). "Histidine Biosynthesis". The Arabidopsis Book. 9: e0141. doi:10.1199/tab.0141. PMC 3266711. PMID 22303266.
- ^ Roberts, John D. (2000). ABCs of FT-NMR. Sausalito, CA: University Science Books. pp. 258–9. ISBN 978-1-891389-18-4.
- ^ Bornhorst, J. A.; Falke, J. J. (2000). "Purification of proteins using polyhistidine affinity tags". Applications of Chimeric Genes and Hybrid Proteins Part A: Gene Expression and Protein Purification. Methods in Enzymology. Vol. 326. pp. 245–254. doi:10.1016/s0076-6879(00)26058-8. ISBN 978-0-12-182227-9. ISSN 0076-6879. PMC 2909483. PMID 11036646.
- ^ Watly, Joanna; Simonovsky, Eyal; Barbosa, Nuno; Spodzieja, Marta; Wieczorek, Robert; Rodziewicz-Motowidlo, Sylwia; Miller, Yifat; Kozlowski, Henryk (2015-08-17). "African Viper Poly-His Tag Peptide Fragment Efficiently Binds Metal Ions and Is Folded into an α-Helical Structure". Inorganic Chemistry. 54 (16): 7692–7702. doi:10.1021/acs.inorgchem.5b01029. ISSN 1520-510X. PMID 26214303.
- ^ Roche Biochemical Pathways Map Roche biochemical pathways map
- ^ a b Alifano, P; Fani, R; Liò, P; Lazcano, A; Bazzicalupo, M; Carlomagno, M S; Bruni, C B (1996-03-01). "Histidine biosynthetic pathway and genes: structure, regulation, and evolution". Microbiological Reviews. 60 (1): 44–69. doi:10.1128/MMBR.60.1.44-69.1996. ISSN 0146-0749. PMC 239417. PMID 8852895.
- ^ a b Kulis-Horn, Robert K; Persicke, Marcus; Kalinowski, Jörn (2014-01-01). "Histidine biosynthesis, its regulation and biotechnological application in Corynebacterium glutamicum". Microbial Biotechnology. 7 (1): 5–25. doi:10.1111/1751-7915.12055. ISSN 1751-7915. PMC 3896937. PMID 23617600.
- ^ Adams, E. (1955-11-01). "L-Histidinal, a biosynthetic precursor of histidine". The Journal of Biological Chemistry. 217 (1): 325–344. doi:10.1016/S0021-9258(19)57184-8. ISSN 0021-9258. PMID 13271397.
- ^ a b Ahmed A. Organization of the histidine-3 region of Neurospora. Mol Gen Genet. 1968;103(2):185-93. doi: 10.1007/BF00427145. PMID 4306011
- ^ DeNofrio, Jan (2011-02-08). "How come plants can make essential amino acids but people can't?". The Tech Interactive. Ask a Geneticist. Retrieved 2024-08-04.
- ^ Stepansky, A.; Leustek, T. (2006-03-01). "Histidine biosynthesis in plants". Amino Acids. 30 (2): 127–142. doi:10.1007/s00726-005-0247-0. ISSN 0939-4451. PMID 16547652. S2CID 23733445.
- ^ Cheng, Yongsong; Zhou, Yunjiao; Yang, Lei; Zhang, Chenglin; Xu, Qingyang; Xie, Xixian; Chen, Ning (2013-05-01). "Modification of histidine biosynthesis pathway genes and the impact on production of L-histidine in Corynebacterium glutamicum". Biotechnology Letters. 35 (5): 735–741. doi:10.1007/s10529-013-1138-1. ISSN 1573-6776. PMID 23355034. S2CID 18380727.
- ^ a b c Board review series (BRS)-- Biochemistry, Molecular Biology, and Genetics (fifth edition): Swanson, Kim, Glucksman
- ^ Coote, J. G.; Hassall, H. (1973-03-01). "The degradation of l-histidine, imidazolyl-l-lactate and imidazolylpropionate by Pseudomonas testosteroni". Biochemical Journal. 132 (3): 409–422. doi:10.1042/bj1320409. ISSN 0264-6021. PMC 1177604. PMID 4146796.
- ^ Mehler, A. H.; Tabor, H. (1953-04-01). "Deamination of histidine to form urocanic acid in liver". The Journal of Biological Chemistry. 201 (2): 775–784. doi:10.1016/S0021-9258(18)66234-9. ISSN 0021-9258. PMID 13061415.
- ^ Andersen, Hjalte H.; Elberling, Jesper; Arendt-Nielsen, Lars (2015-09-01). "Human surrogate models of histaminergic and non-histaminergic itch" (PDF). Acta Dermato-Venereologica. 95 (7): 771–777. doi:10.2340/00015555-2146. ISSN 1651-2057. PMID 26015312.
- ^ "3-Methylhistidine". HMDB Version 4.0. Human Metabolome Database. 20 December 2017. Retrieved 25 December 2017.
- ^ Derave, Wim; Everaert, Inge; Beeckman, Sam; Baguet, Audrey (2010-03-01). "Muscle carnosine metabolism and beta-alanine supplementation in relation to exercise and training". Sports Medicine. 40 (3): 247–263. doi:10.2165/11530310-000000000-00000. hdl:1854/LU-897781. ISSN 1179-2035. PMID 20199122. S2CID 7661250.
- ^ Fahey, Robert C. (2001). "Novelthiols Ofprokaryotes". Annual Review of Microbiology. 55: 333–56. doi:10.1146/annurev.micro.55.1.333. PMID 11544359.
- ^ Institute of Medicine (2002). "Protein and Amino Acids". Dietary Reference Intakes for Energy, Carbohydrates, Fiber, Fat, Fatty Acids, Cholesterol, Protein, and Amino Acids. Washington, DC: The National Academies Press. pp. 589–768. doi:10.17226/10490. ISBN 978-0-309-08525-0.
- ^ Holeček, Milan (2020-03-22). "Histidine in Health and Disease: Metabolism, Physiological Importance, and Use as a Supplement". Nutrients. 12 (3): 848. doi:10.3390/nu12030848. ISSN 2072-6643. PMC 7146355. PMID 32235743.