Primase
| Toprim domain | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Identifiers | |||||||||
| Symbol | Toprim | ||||||||
| Pfam | PF01751 | ||||||||
| Pfam clan | Toprim-like | ||||||||
| InterPro | IPR006171 | ||||||||
| SCOP2 | 2fcj / SCOPe / SUPFAM | ||||||||
| |||||||||
| Toprim catalytic core | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Identifiers | |||||||||
| Symbol | Toprim_N | ||||||||
| Pfam | PF08275 | ||||||||
| InterPro | IPR013264 | ||||||||
| SCOP2 | 1dd9 / SCOPe / SUPFAM | ||||||||
| |||||||||
| AEP DNA primase, small subunit | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Identifiers | |||||||||
| Symbol | DNA_primase_S | ||||||||
| Pfam | PF01896 | ||||||||
| Pfam clan | AEP | ||||||||
| InterPro | IPR002755 | ||||||||
| SCOP2 | 1g71 / SCOPe / SUPFAM | ||||||||
| |||||||||
| AEP DNA primase, large subunit | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Identifiers | |||||||||
| Symbol | DNA_primase_lrg | ||||||||
| Pfam | PF04104 | ||||||||
| Pfam clan | CL0242 | ||||||||
| InterPro | IPR007238 | ||||||||
| SCOP2 | 1zt2 / SCOPe / SUPFAM | ||||||||
| |||||||||
DNA primase is an enzyme involved in the replication of DNA and is a type of RNA polymerase. Primase catalyzes the synthesis of a short RNA (or DNA in some living organisms[1]) segment called a primer complementary to a ssDNA (single-stranded DNA) template. After this elongation, the RNA piece is removed by a 5' to 3' exonuclease and refilled with DNA.
Function
[edit]
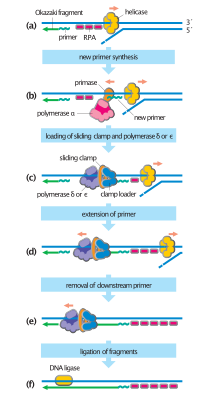
In bacteria, primase binds to the DNA helicase forming a complex called the primosome. Primase is activated by the helicase where it then synthesizes a short RNA primer approximately 11 ±1 nucleotides long, to which new nucleotides can be added by DNA polymerase. Archaeal and eukaryote primases are heterodimeric proteins with one large regulatory and one minuscule catalytic subunit.[2]
The RNA segments are first synthesized by primase and then elongated by DNA polymerase.[3] Then the DNA polymerase forms a protein complex with two primase subunits to form the alpha DNA Polymerase primase complex. Primase is one of the most error prone and slow polymerases.[3] Primases in organisms such as E. coli synthesize around 2000 to 3000 primers at the rate of one primer per second.[4] Primase also acts as a halting mechanism to prevent the leading strand from outpacing the lagging strand by halting the progression of the replication fork.[5] The rate determining step in primase is when the first phosphodiester bond is formed between two molecules of RNA.[3]
The replication mechanisms differ between different bacteria and viruses where the primase covalently link to helicase in viruses such as the T7 bacteriophage.[5] In viruses such as the herpes simplex virus (HSV-1), primase can form complexes with helicase.[6] The primase-helicase complex is used to unwind dsDNA (double-stranded) and synthesizes the lagging strand using RNA primers[6] The majority of primers synthesized by primase are two to three nucleotides long.[6]
Types
[edit]There are two main types of primase: DnaG found in most bacteria, and the AEP (Archaeo-Eukaryote Primase) superfamily found in archaean and eukaryotic primases. While bacterial primases (DnaG-type) are composed of a single protein unit (a monomer) and synthesize RNA primers, AEP primases are usually composed of two different primase units (a heterodimer) and synthesize two-part primers with both RNA and DNA components.[7] While functionally similar, the two primase superfamilies evolved independently of each other.
DnaG
[edit]The crystal structure of primase in E. coli with a core containing the DnaG protein was determined in the year 2000.[4] The DnaG and primase complex is cashew shaped and contains three subdomains.[4] The central subdomain forms a toprim fold which is made of a mixture five beta sheets and six alpha helices.[4][8] The toprim fold is used for binding regulators and metals. The primase uses a phosphotransfer domain for the transfer coordination of metals, which makes it distinct from other polymerases.[4] The side subunits contain a NH2 and COOH terminal made of alpha helixes and beta sheets.[4] The NH2 terminal interacts with a zinc binding domain and COOH-terminal region which interacts with DnaB-ID.[4]
The Toprim fold is also found in topoisomerase and mitochrondrial Twinkle primase/helicase.[8] Some DnaG-like (bacteria-like; InterPro: IPR020607) primases have been found in archaeal genomes.[9]
AEP
[edit]Eukaryote and archaeal primases tend to be more similar to each other, in terms of structure and mechanism, than they are to bacterial primases.[10][11] The archaea-eukaryotic primase (AEP) superfamily, which most eukaryal and archaeal primase catalytic subunits belong to, has recently been redefined as a primase-polymerase family in recognition of the many other roles played by enzymes in this family.[12] This classification also emphasizes the broad origins of AEP primases; the superfamily is now recognized as transitioning between RNA and DNA functions.[13]
Archaeal and eukaryote primases are heterodimeric proteins with one large regulatory (human PRIM2, p58) and one small catalytic subunit (human PRIM1, p48/p49).[2] The large subunit contains a N-terminal 4Fe–4S cluster, split out in some archaea as PriX/PriCT.[14] The large subunit is implicated in improving the activity and specificity of the small subunit. For example, removing the part corresponding to the large subunit in a fusion protein PolpTN2 results in a slower enzyme with reverse transcriptase activity.[13]
Multifunctional primases
[edit]
The AEP family of primase-polymerases has diverse features beyond making only primers. In addition to priming DNA during replication, AEP enzymes may have additional functions in the DNA replication process, such as polymerization of DNA or RNA, terminal transfer, translesion synthesis (TLS), non-homologous end joining (NHEJ),[12] and possibly in restarting stalled replication forks.[15] Primases typically synthesize primers from ribonucleotides (NTPs); however, primases with polymerase capabilities also have an affinity for deoxyribonucleotides (dNTPs).[16][11] Primases with terminal transferase functionality are capable of adding nucleotides to the 3’ end of a DNA strand independently of a template. Other enzymes involved in DNA replication, such as helicases, may also exhibit primase activity.[17]
In eukaryotes and archaea
[edit]Human PrimPol (ccdc111[16]) serves both primase and polymerase functions, like many archaeal primases; exhibits terminal transferase activity in the presence of manganese; and plays a significant role in translesion synthesis[18] and in restarting stalled replication forks. PrimPol is actively recruited to damaged sites through its interaction with RPA, an adapter protein that facilitates DNA replication and repair.[15] PrimPol has a zinc finger domain similar to that of some viral primases, which is essential for translesion synthesis and primase activity and may regulate primer length.[18] Unlike most primases, PrimPol is uniquely capable of starting DNA chains with dNTPs.[16]
PriS, the archaeal primase small subunit, has a role in translesion synthesis (TLS) and can bypass common DNA lesions. Most archaea lack the specialized polymerases that perform TLS in eukaryotes and bacteria.[19] PriS alone preferentially synthesizes strings of DNA; but in combination with PriL, the large subunit, RNA polymerase activity is increased.[20]
In Sulfolobus solfataricus, the primase heterodimer PriSL can act as a primase, polymerase, and terminal transferase. PriSL is thought to initiate primer synthesis with NTPs and then switch to dNTPs. The enzyme can polymerize RNA or DNA chains, with DNA products reaching as long as 7000 nucleotides (7 kb). It is suggested that this dual functionality may be a common feature of archaeal primases.[11]
In bacteria
[edit]AEP multifutional primases also appear in bacteria and phages that infect them. They can display novel domain organizations with domains that bring even more functions beyond polymerization.[14]
Bacterial LigD (A0R3R7) is primarily involved in the NHEJ pathway. It has an AEP superfamily polymerase/primase domain, a 3'-phosphoesterase domain, and a ligase domain. It is also capable of primase, DNA and RNA polymerase, and terminal transferase activity. DNA polymerization activity can produce chains over 7000 nucleotides (7 kb) in length, while RNA polymerization produces chains up to 1 kb long.[21]
In viruses and plasmids
[edit]AEP enzymes are widespread, and can be found encoded in mobile genetic elements including virus/phages and plasmids. They either use them as a sole replication protein or in combination with other replication-associated proteins, such as helicases and, less frequently, DNA polymerases.[22] Whereas the presence of AEP in eukaryotic and archaeal viruses is expected in that they mirror their hosts,[22] bacterial viruses and plasmids also as frequently encode AEP-superfamily enzymes as they do DnaG-family primases.[14] A great diversity of AEP families has been uncovered in various bacterial plasmids by comparative genomics surveys.[14] Their evolutionary history is currently unknown, as these found in bacteria and bacteriophages appear too different from their archaeo-eukaryotic homologs for a recent horizontal gene transfer.[22]
MCM-like helicase in Bacillus cereus strain ATCC 14579 (BcMCM; Q81EV1) is an SF6 helicase fused with an AEP primase. The enzyme has both primase and polymerase functions in addition to helicase function. The gene coding for it is found in a prophage.[17] It bears homology to ORF904 of plasmid pRN1 from Sulfolobus islandicus, which has an AEP PrimPol domain.[23] Vaccinia virus D5 and HSV Primase are examples of AEP-helicase fusion as well.[12][6]
PolpTN2 is an Archaeal primase found in the TN2 plasmid. A fusion of domains homologous to PriS and PriL, it exhibits both primase and DNA polymerase activity, as well as terminal transferase function. Unlike most primases, PolpTN2 forms primers composed exclusively of dNTPs.[13] Unexpectedly, when the PriL-like domain was truncated, PolpTN2 could also synthesize DNA on the RNA template, i.e., acted as an RNA-dependent DNA polymerase (reverse transcriptase).[13]
Even DnaG primases can have extra functions, if given the right domains. The T7 phage gp4 is a DnaG primase-helicase fusion, and performs both functions in replication.[5]
References
[edit]- ^ Bocquier AA, Liu L, Cann IK, Komori K, Kohda D, Ishino Y (March 2001). "Archaeal primase: bridging the gap between RNA and DNA polymerases". Current Biology. 11 (6): 452–6. doi:10.1016/s0960-9822(01)00119-1. PMID 11301257.
- ^ a b Baranovskiy AG, Zhang Y, Suwa Y, Babayeva ND, Gu J, Pavlov YI, Tahirov TH (February 2015). "Crystal structure of the human primase". The Journal of Biological Chemistry. 290 (9): 5635–46. doi:10.1074/jbc.M114.624742. PMC 4342476. PMID 25550159.
- ^ a b c Griep MA (August 1995). "Primase structure and function". Indian Journal of Biochemistry & Biophysics. 32 (4): 171–8. PMID 8655184.
- ^ a b c d e f g Keck JL, Roche DD, Lynch AS, Berger JM (March 2000). "Structure of the RNA polymerase domain of E. coli primase". Science. 287 (5462): 2482–6. Bibcode:2000Sci...287.2482K. doi:10.1126/science.287.5462.2482. PMID 10741967.
- ^ a b c Lee JB, Hite RK, Hamdan SM, Xie XS, Richardson CC, van Oijen AM (February 2006). "DNA primase acts as a molecular brake in DNA replication" (PDF). Nature. 439 (7076): 621–4. Bibcode:2006Natur.439..621L. doi:10.1038/nature04317. PMID 16452983. S2CID 3099842.
- ^ a b c d Cavanaugh NA, Kuchta RD (January 2009). "Initiation of new DNA strands by the herpes simplex virus-1 primase-helicase complex and either herpes DNA polymerase or human DNA polymerase alpha". The Journal of Biological Chemistry. 284 (3): 1523–32. doi:10.1074/jbc.M805476200. PMC 2615532. PMID 19028696.
- ^ Keck JL, Berger JM (January 2001). "Primus inter pares (first among equals)". Nature Structural Biology. 8 (1): 2–4. doi:10.1038/82996. PMID 11135655. S2CID 17108681.
- ^ a b Aravind L, Leipe DD, Koonin EV (September 1998). "Toprim--a conserved catalytic domain in type IA and II topoisomerases, DnaG-type primases, OLD family nucleases and RecR proteins". Nucleic Acids Research. 26 (18): 4205–13. doi:10.1093/nar/26.18.4205. PMC 147817. PMID 9722641.
- ^ Hou L, Klug G, Evguenieva-Hackenberg E (March 2013). "The archaeal DnaG protein needs Csl4 for binding to the exosome and enhances its interaction with adenine-rich RNAs". RNA Biology. 10 (3): 415–24. doi:10.4161/rna.23450. PMC 3672285. PMID 23324612.
- ^ Iyer LM, Koonin EV, Leipe DD, Aravind L (2005). "Origin and evolution of the archaeo-eukaryotic primase superfamily and related palm-domain proteins: structural insights and new members". Nucleic Acids Research. 33 (12): 3875–96. doi:10.1093/nar/gki702. PMC 1176014. PMID 16027112.
- ^ a b c Lao-Sirieix SH, Bell SD (December 2004). "The heterodimeric primase of the hyperthermophilic archaeon Sulfolobus solfataricus possesses DNA and RNA primase, polymerase and 3'-terminal nucleotidyl transferase activities". Journal of Molecular Biology. 344 (5): 1251–63. doi:10.1016/j.jmb.2004.10.018. PMID 15561142.
- ^ a b c d Guilliam TA, Keen BA, Brissett NC, Doherty AJ (August 2015). "Primase-polymerases are a functionally diverse superfamily of replication and repair enzymes". Nucleic Acids Research. 43 (14): 6651–64. doi:10.1093/nar/gkv625. PMC 4538821. PMID 26109351.
- ^ a b c d Gill S, Krupovic M, Desnoues N, Béguin P, Sezonov G, Forterre P (April 2014). "A highly divergent archaeo-eukaryotic primase from the Thermococcus nautilus plasmid, pTN2". Nucleic Acids Research. 42 (6): 3707–19. doi:10.1093/nar/gkt1385. PMC 3973330. PMID 24445805.
- ^ a b c d Kazlauskas D, Sezonov G, Charpin N, Venclovas Č, Forterre P, Krupovic M (March 2018). "Novel Families of Archaeo-Eukaryotic Primases Associated with Mobile Genetic Elements of Bacteria and Archaea". Journal of Molecular Biology. 430 (5): 737–750. doi:10.1016/j.jmb.2017.11.014. PMC 5862659. PMID 29198957.
- ^ a b Wan L, Lou J, Xia Y, Su B, Liu T, Cui J, Sun Y, Lou H, Huang J (December 2013). "hPrimpol1/CCDC111 is a human DNA primase-polymerase required for the maintenance of genome integrity". EMBO Reports. 14 (12): 1104–12. doi:10.1038/embor.2013.159. PMC 3981091. PMID 24126761.
- ^ a b c García-Gómez S, Reyes A, Martínez-Jiménez MI, Chocrón ES, Mourón S, Terrados G, Powell C, Salido E, Méndez J, Holt IJ, Blanco L (November 2013). "PrimPol, an archaic primase/polymerase operating in human cells". Molecular Cell. 52 (4): 541–53. doi:10.1016/j.molcel.2013.09.025. PMC 3899013. PMID 24207056.
- ^ a b Sanchez-Berrondo J, Mesa P, Ibarra A, Martínez-Jiménez MI, Blanco L, Méndez J, Boskovic J, Montoya G (February 2012). "Molecular architecture of a multifunctional MCM complex". Nucleic Acids Research. 40 (3): 1366–80. doi:10.1093/nar/gkr831. PMC 3273815. PMID 21984415.
- ^ a b Keen BA, Jozwiakowski SK, Bailey LJ, Bianchi J, Doherty AJ (May 2014). "Molecular dissection of the domain architecture and catalytic activities of human PrimPol". Nucleic Acids Research. 42 (9): 5830–45. doi:10.1093/nar/gku214. PMC 4027207. PMID 24682820.
- ^ Jozwiakowski SK, Borazjani Gholami F, Doherty AJ (February 2015). "Archaeal replicative primases can perform translesion DNA synthesis". Proceedings of the National Academy of Sciences of the United States of America. 112 (7): E633-8. Bibcode:2015PNAS..112E.633J. doi:10.1073/pnas.1412982112. PMC 4343091. PMID 25646444.
- ^ Barry ER, Bell SD (December 2006). "DNA replication in the archaea". Microbiology and Molecular Biology Reviews. 70 (4): 876–87. doi:10.1128/MMBR.00029-06. PMC 1698513. PMID 17158702.
- ^ Lao-Sirieix SH, Pellegrini L, Bell SD (October 2005). "The promiscuous primase". Trends in Genetics. 21 (10): 568–72. doi:10.1016/j.tig.2005.07.010. PMID 16095750.
- ^ a b c Kazlauskas D, Krupovic M, Venclovas Č (June 2016). "The logic of DNA replication in double-stranded DNA viruses: insights from global analysis of viral genomes". Nucleic Acids Research. 44 (10): 4551–64. doi:10.1093/nar/gkw322. PMC 4889955. PMID 27112572.
- ^ Lipps G, Weinzierl AO, von Scheven G, Buchen C, Cramer P (February 2004). "Structure of a bifunctional DNA primase-polymerase". Nature Structural & Molecular Biology. 11 (2): 157–62. doi:10.1038/nsmb723. PMID 14730355. S2CID 25123984.
