RAB11B
Ras-related protein Rab-11B is a protein that in humans is encoded by the RAB11B gene.[5][6] Rab11b is reported as most abundantly expressed in brain, heart and testes.
Rab (Ras-related in brain) proteins form the largest section of the Ras superfamily of small GTPases. The Rab family proteins regulate intracellular membrane trafficking processes including vesicle budding, tethering, and fusion. The isoforms Rab11a, Rab11b, and Rab11c/Rab25 constitute the Rab11 subfamily based on specific sequence motifs.[7] While RAB11A is located on chromosome 15[8] and RAB11C on chromosome 1, RAB11B is placed on chromosome 19. Rab11 proteins are implicated in endocytosis and exocytosis.[9] Rab11b is reported as most abundantly expressed in brain, heart and testes.[10] Early studies with deletions of RAB11 homologs in Saccharomyces cerevisiae proved their importance in cell survival.[11][12] Despite sharing high sequence homology, Rab11a and Rab11b appear to reside within distinct vesicle compartments.[13] Majority of Rab11b neither colocalize with transferrin receptor nor with the polymeric IgA receptor. This protein also exhibits a dependence on the microtubule cytoskeleton that is different from Rab11a.[13] High sequence diversity in the C-terminal hypervariable region is responsible for variable membrane targeting between these proteins.
Function
[edit]Members of the Rab11 subfamily act in recycling of proteins from the endosomes to the plasma membrane, in transport of molecules from the trans-Golgi network to the plasma membrane and in phagocytosis. This subfamily also acts in polarized transport in epithelial cells.[14][15][16][17][18] Whereas most studies refer to the Rab11a isoform, little is known about Rab11b so far. Rab11b localizes predominantly in the pericentriolar recycling compartment and serves as an important component of the vesicular machinery.[19] It is required for the transfer of internalized transferrin from the recycling compartment to the plasma membrane for which active Rab11b as well as GTP hydrolysis is necessary.[19]
Structure
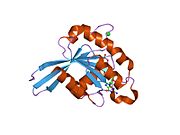
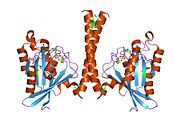
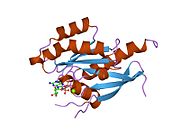
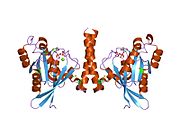
[edit]All Ras GTPases consist of a similar core structure and highly conserved P-loop, switch 1 and switch 2 regions. The Rab11b monomer exhibits a typical Ras-like, small GTPase fold with a six stranded β-sheet core (β1-β6) surrounded by five major α-helices (α1-α5)[16] and one minor α-helix (α6). According to the sequence similarity to other Rab GTPases can be assumed that they show closely resembling characteristics in nucleotide binding and hydrolysis. However, Rab11 isoforms could differ in hydrolysis kinetics owing to the differences in conformation, since Rab11a and Rab11b do not show an α-helical switch 2 region like other Rab GTPases. Rab11b shares 90% amino acid identity to Rab11a.[16] Kinetic experiments with Rab11a/b and Rab11-interacting proteins (FIPs) indicate that FIPs cannot differentiate between GTP-bound Rab11a and Rab11b in vitro.[20] The major divergence reveals in the inactive state. While Pasqualato et al. crystallized inactive Rab11a as a dimer in the asymmetric unit, Scapin et al. observed single crystallographically independent monomers of both the GDP- and the GppNHp-bound Rab11b structures.[16][21]


Clinical significance
[edit]Due to their crucial importance in vesicle transport and recycling, Rab11 proteins are linked to various non-pathogen or pathogen induced diseases. Most of the published data do not specify whether it is the a- or the b-isoform. Rab11 proteins have been implicated in Alzheimer's disease,[22][23] Arthrogryposis-renal dysfunction-cholestasis (ARC),[24] Batten disease,[25] and Charcot-Marie-Tooth Neuropathy Type 4C (CMT4C).[26] Intracellular bacteria Chlamydia pneumoniae and Chlamydia trachomatis that replicate in membrane bound compartments hijack the trafficking machinery recruiting Rab GTPases to promote their replication within the host cell. Knock down of Rab11 decreased the formation of infectious particles.[27][28][29] Recent studies report a similar use of intracellular trafficking by Hantavirus and Influenza A virus. Replicated viruses benefit from Rab11 mediated recycling endosome pathway to exit the cell and infect surrounding tissue.[30][31][32][33]
References
[edit]- ^ a b c GRCh38: Ensembl release 89: ENSG00000185236 – Ensembl, May 2017
- ^ a b c GRCm38: Ensembl release 89: ENSMUSG00000077450 – Ensembl, May 2017
- ^ "Human PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- ^ "Mouse PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- ^ Zhu AX, Zhao Y, Flier JS (Dec 1994). "Molecular cloning of two small GTP-binding proteins from human skeletal muscle". Biochemical and Biophysical Research Communications. 205 (3): 1875–82. doi:10.1006/bbrc.1994.2889. PMID 7811277.
- ^ "Entrez Gene: RAB11B RAB11B, member RAS oncogene family".
- ^ Bhartur SG, Calhoun BC, Woodrum J, Kurkjian J, Iyer S, Lai F, Goldenring JR (Mar 2000). "Genomic structure of murine Rab11 family members". Biochemical and Biophysical Research Communications. 269 (2): 611–7. doi:10.1006/bbrc.2000.2334. PMID 10708602.
- ^ Gromov PS, Celis JE, Hansen C, Tommerup N, Gromova I, Madsen P (Jun 1998). "Human rab11a: transcription, chromosome mapping and effect on the expression levels of host GTP-binding proteins". FEBS Letters. 429 (3): 359–64. doi:10.1016/s0014-5793(98)00607-3. PMID 9662449. S2CID 22139183.
- ^ Wilcke M, Johannes L, Galli T, Mayau V, Goud B, Salamero J (Dec 2000). "Rab11 regulates the compartmentalization of early endosomes required for efficient transport from early endosomes to the trans-golgi network". The Journal of Cell Biology. 151 (6): 1207–20. doi:10.1083/jcb.151.6.1207. PMC 2190589. PMID 11121436.
- ^ Lai F, Stubbs L, Artzt K (Aug 1994). "Molecular analysis of mouse Rab11b: a new type of mammalian YPT/Rab protein". Genomics. 22 (3): 610–6. doi:10.1006/geno.1994.1434. PMID 8001972.
- ^ Benli M, Döring F, Robinson DG, Yang X, Gallwitz D (Dec 1996). "Two GTPase isoforms, Ypt31p and Ypt32p, are essential for Golgi function in yeast". The EMBO Journal. 15 (23): 6460–75. doi:10.1002/j.1460-2075.1996.tb01037.x. PMC 452471. PMID 8978673.
- ^ Jedd G, Mulholland J, Segev N (May 1997). "Two new Ypt GTPases are required for exit from the yeast trans-Golgi compartment". The Journal of Cell Biology. 137 (3): 563–80. doi:10.1083/jcb.137.3.563. PMC 2139891. PMID 9151665.
- ^ Chen W, Feng Y, Chen D, Wandinger-Ness A (Nov 1998). "Rab11 is required for trans-golgi network-to-plasma membrane transport and a preferential target for GDP dissociation inhibitor". Molecular Biology of the Cell. 9 (11): 3241–57. doi:10.1091/mbc.9.11.3241. PMC 25617. PMID 9802909.
- ^ Cox D, Lee DJ, Dale BM, Calafat J, Greenberg S (Jan 2000). "A Rab11-containing rapidly recycling compartment in macrophages that promotes phagocytosis". Proceedings of the National Academy of Sciences of the United States of America. 97 (2): 680–5. Bibcode:2000PNAS...97..680C. doi:10.1073/pnas.97.2.680. PMC 15390. PMID 10639139.
- ^ a b c d e f Scapin SM, Carneiro FR, Alves AC, Medrano FJ, Guimarães BG, Zanchin NI (Jun 2006). "The crystal structure of the small GTPase Rab11b reveals critical differences relative to the Rab11a isoform". Journal of Structural Biology. 154 (3): 260–8. doi:10.1016/j.jsb.2006.01.007. PMID 16545962.
- ^ Ullrich O, Reinsch S, Urbé S, Zerial M, Parton RG (Nov 1996). "Rab11 regulates recycling through the pericentriolar recycling endosome". The Journal of Cell Biology. 135 (4): 913–24. doi:10.1083/jcb.135.4.913. PMC 2133374. PMID 8922376.
- ^ Wang X, Kumar R, Navarre J, Casanova JE, Goldenring JR (Sep 2000). "Regulation of vesicle trafficking in madin-darby canine kidney cells by Rab11a and Rab25". The Journal of Biological Chemistry. 275 (37): 29138–46. doi:10.1074/jbc.M004410200. PMID 10869360.
- ^ a b Schlierf B, Fey GH, Hauber J, Hocke GM, Rosorius O (Aug 2000). "Rab11b is essential for recycling of transferrin to the plasma membrane". Experimental Cell Research. 259 (1): 257–65. doi:10.1006/excr.2000.4947. PMID 10942597.
- ^ Shiba T, Koga H, Shin HW, Kawasaki M, Kato R, Nakayama K, Wakatsuki S (Oct 2006). "Structural basis for Rab11-dependent membrane recruitment of a family of Rab11-interacting protein 3 (FIP3)/Arfophilin-1". Proceedings of the National Academy of Sciences of the United States of America. 103 (42): 15416–21. Bibcode:2006PNAS..10315416S. doi:10.1073/pnas.0605357103. PMC 1622838. PMID 17030804.
- ^ Pasqualato S, Senic-Matuglia F, Renault L, Goud B, Salamero J, Cherfils J (Mar 2004). "The structural GDP/GTP cycle of Rab11 reveals a novel interface involved in the dynamics of recycling endosomes". The Journal of Biological Chemistry. 279 (12): 11480–8. doi:10.1074/jbc.M310558200. PMID 14699104.
- ^ Greenfield JP, Leung LW, Cai D, Kaasik K, Gross RS, Rodriguez-Boulan E, Greengard P, Xu H (Apr 2002). "Estrogen lowers Alzheimer beta-amyloid generation by stimulating trans-Golgi network vesicle biogenesis". The Journal of Biological Chemistry. 277 (14): 12128–36. doi:10.1074/jbc.M110009200. PMID 11823458.
- ^ Dumanchin C, Czech C, Campion D, Cuif MH, Poyot T, Martin C, Charbonnier F, Goud B, Pradier L, Frebourg T (Jul 1999). "Presenilins interact with Rab11, a small GTPase involved in the regulation of vesicular transport". Human Molecular Genetics. 8 (7): 1263–9. doi:10.1093/hmg/8.7.1263. PMID 10369872.
- ^ Cullinane AR, Straatman-Iwanowska A, Zaucker A, Wakabayashi Y, Bruce CK, Luo G, Rahman F, Gürakan F, Utine E, Ozkan TB, Denecke J, Vukovic J, Di Rocco M, Mandel H, Cangul H, Matthews RP, Thomas SG, Rappoport JZ, Arias IM, Wolburg H, Knisely AS, Kelly DA, Müller F, Maher ER, Gissen P (Apr 2010). "Mutations in VIPAR cause an arthrogryposis, renal dysfunction and cholestasis syndrome phenotype with defects in epithelial polarization". Nature Genetics. 42 (4): 303–12. doi:10.1038/ng.538. PMC 5308204. PMID 20190753.
- ^ Luiro K, Yliannala K, Ahtiainen L, Maunu H, Järvelä I, Kyttälä A, Jalanko A (Dec 2004). "Interconnections of CLN3, Hook1 and Rab proteins link Batten disease to defects in the endocytic pathway". Human Molecular Genetics. 13 (23): 3017–27. doi:10.1093/hmg/ddh321. PMID 15471887.
- ^ Stendel C, Roos A, Kleine H, Arnaud E, Ozçelik M, Sidiropoulos PN, Zenker J, Schüpfer F, Lehmann U, Sobota RM, Litchfield DW, Lüscher B, Chrast R, Suter U, Senderek J (Aug 2010). "SH3TC2, a protein mutant in Charcot-Marie-Tooth neuropathy, links peripheral nerve myelination to endosomal recycling". Brain. 133 (Pt 8): 2462–74. doi:10.1093/brain/awq168. hdl:20.500.11850/157734. PMID 20826437.
- ^ Cortes C, Rzomp KA, Tvinnereim A, Scidmore MA, Wizel B (Dec 2007). "Chlamydia pneumoniae inclusion membrane protein Cpn0585 interacts with multiple Rab GTPases". Infection and Immunity. 75 (12): 5586–96. doi:10.1128/IAI.01020-07. PMC 2168330. PMID 17908815.
- ^ Rejman Lipinski A, Heymann J, Meissner C, Karlas A, Brinkmann V, Meyer TF, Heuer D (Oct 2009). "Rab6 and Rab11 regulate Chlamydia trachomatis development and golgin-84-dependent Golgi fragmentation". PLOS Pathogens. 5 (10): e1000615. doi:10.1371/journal.ppat.1000615. PMC 2752117. PMID 19816566.
- ^ Rzomp KA, Scholtes LD, Briggs BJ, Whittaker GR, Scidmore MA (Oct 2003). "Rab GTPases are recruited to chlamydial inclusions in both a species-dependent and species-independent manner". Infection and Immunity. 71 (10): 5855–70. doi:10.1128/IAI.71.10.5855-5870.2003. PMC 201052. PMID 14500507.
- ^ Amorim MJ, Bruce EA, Read EK, Foeglein A, Mahen R, Stuart AD, Digard P (May 2011). "A Rab11- and microtubule-dependent mechanism for cytoplasmic transport of influenza A virus viral RNA". Journal of Virology. 85 (9): 4143–56. doi:10.1128/JVI.02606-10. PMC 3126276. PMID 21307188.
- ^ Bruce EA, Digard P, Stuart AD (Jun 2010). "The Rab11 pathway is required for influenza A virus budding and filament formation". Journal of Virology. 84 (12): 5848–59. doi:10.1128/JVI.00307-10. PMC 2876627. PMID 20357086.
- ^ Momose F, Sekimoto T, Ohkura T, Jo S, Kawaguchi A, Nagata K, Morikawa Y (2011-06-22). "Apical transport of influenza A virus ribonucleoprotein requires Rab11-positive recycling endosome". PLOS ONE. 6 (6): e21123. Bibcode:2011PLoSO...621123M. doi:10.1371/journal.pone.0021123. PMC 3120830. PMID 21731653.
- ^ Rowe, Regina K.; Jason W. Suszko; Andrew Pekosz (2008-12-20). "Roles for the recycling endosome, Rab8, and Rab11 in hantavirus release from epithelial cells". Virology. 382 (2): 239–249. doi:10.1016/j.virol.2008.09.021. ISSN 0042-6822. PMC 2648827. PMID 18951604.
Further reading
[edit]- Maruyama K, Sugano S (Jan 1994). "Oligo-capping: a simple method to replace the cap structure of eukaryotic mRNAs with oligoribonucleotides". Gene. 138 (1–2): 171–4. doi:10.1016/0378-1119(94)90802-8. PMID 8125298.
- Suzuki Y, Yoshitomo-Nakagawa K, Maruyama K, Suyama A, Sugano S (Oct 1997). "Construction and characterization of a full length-enriched and a 5'-end-enriched cDNA library". Gene. 200 (1–2): 149–56. doi:10.1016/S0378-1119(97)00411-3. PMID 9373149.
- Schlierf B, Fey GH, Hauber J, Hocke GM, Rosorius O (Aug 2000). "Rab11b is essential for recycling of transferrin to the plasma membrane". Experimental Cell Research. 259 (1): 257–65. doi:10.1006/excr.2000.4947. PMID 10942597.
- Lapierre LA, Kumar R, Hales CM, Navarre J, Bhartur SG, Burnette JO, Provance DW, Mercer JA, Bähler M, Goldenring JR (Jun 2001). "Myosin vb is associated with plasma membrane recycling systems". Molecular Biology of the Cell. 12 (6): 1843–57. doi:10.1091/mbc.12.6.1843. PMC 37346. PMID 11408590.
- Prekeris R, Davies JM, Scheller RH (Oct 2001). "Identification of a novel Rab11/25 binding domain present in Eferin and Rip proteins". The Journal of Biological Chemistry. 276 (42): 38966–70. doi:10.1074/jbc.M106133200. PMID 11481332.
- Hales CM, Griner R, Hobdy-Henderson KC, Dorn MC, Hardy D, Kumar R, Navarre J, Chan EK, Lapierre LA, Goldenring JR (Oct 2001). "Identification and characterization of a family of Rab11-interacting proteins". The Journal of Biological Chemistry. 276 (42): 39067–75. doi:10.1074/jbc.M104831200. PMID 11495908.
- Khvotchev MV, Ren M, Takamori S, Jahn R, Südhof TC (Nov 2003). "Divergent functions of neuronal Rab11b in Ca2+-regulated versus constitutive exocytosis". The Journal of Neuroscience. 23 (33): 10531–9. doi:10.1523/JNEUROSCI.23-33-10531.2003. PMC 6740915. PMID 14627637.
- Scapin SM, Carneiro FR, Alves AC, Medrano FJ, Guimarães BG, Zanchin NI (Jun 2006). "The crystal structure of the small GTPase Rab11b reveals critical differences relative to the Rab11a isoform". Journal of Structural Biology. 154 (3): 260–8. doi:10.1016/j.jsb.2006.01.007. PMID 16545962.