Purine nucleoside phosphorylase
This article includes a list of general references, but it lacks sufficient corresponding inline citations. (September 2017) |
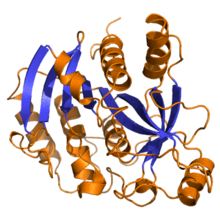
| purine-nucleoside phosphorylase | |||||||||
|---|---|---|---|---|---|---|---|---|---|
 | |||||||||
| Identifiers | |||||||||
| EC no. | 2.4.2.1 | ||||||||
| CAS no. | 9030-21-1 | ||||||||
| Databases | |||||||||
| IntEnz | IntEnz view | ||||||||
| BRENDA | BRENDA entry | ||||||||
| ExPASy | NiceZyme view | ||||||||
| KEGG | KEGG entry | ||||||||
| MetaCyc | metabolic pathway | ||||||||
| PRIAM | profile | ||||||||
| PDB structures | RCSB PDB PDBe PDBsum | ||||||||
| Gene Ontology | AmiGO / QuickGO | ||||||||
| |||||||||
| Purine nucleoside phosphorylase | |||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Identifiers | |||||||||||||||||||||||||||||||||||||||||||||||||||
| Aliases | Purine_phosphorylaseIPR011268 | ||||||||||||||||||||||||||||||||||||||||||||||||||
| External IDs | GeneCards: [1]; OMA:- orthologs | ||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
| Wikidata | |||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
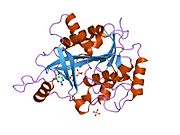
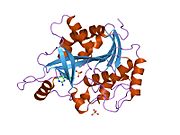
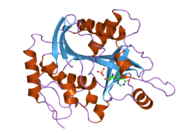
Purine nucleoside phosphorylase, PNP, PNPase or inosine phosphorylase (EC 2.4.2.1) is an enzyme that in humans is encoded by the NP gene.[2] It catalyzes the chemical reaction
- purine nucleoside + phosphate purine + alpha-D-ribose 1-phosphate
Thus, the two substrates of this enzyme are a purine nucleoside and phosphate, whereas its products are a purine and alpha-D-ribose 1-phosphate.
Nomenclature
[edit]This enzyme belongs to the family of glycosyltransferases, specifically the pentosyltransferases. The systematic name of this enzyme class is purine-nucleoside:phosphate ribosyltransferase.
Other names in common use include:
- inosine phosphorylase
- PNPase
- PUNPI
- PUNPII
- inosine-guanosine phosphorylase
- nucleotide phosphatase
- purine deoxynucleoside phosphorylase
- purine deoxyribonucleoside phosphorylase
- purine nucleoside phosphorylase
- purine ribonucleoside phosphorylas
This enzyme participates in 3 metabolic pathways: purine metabolism, pyrimidine metabolism, and nicotinate and nicotinamide metabolism.
Function
[edit]Purine nucleoside phosphorylase is an enzyme involved in purine metabolism. PNP metabolizes inosine into hypoxanthine and guanosine into guanine, in each case creating ribose phosphate. Note: adenosine is first metabolized to inosine via the enzyme adenosine deaminase.[3]

Nucleoside phosphorylase is an enzyme which cleaves a nucleoside by phosphorylating the ribose to produce a nucleobase and ribose 1 phosphate. It is one enzyme of the nucleotide salvage pathways. These pathways allow the cell to produce nucleotide monophosphates when the de novo synthesis pathway has been interrupted or is non-existent (as is the case in the brain). Often the de novo pathway is interrupted as a result of chemotherapy drugs such as methotrexate or aminopterin.
All salvage pathway enzymes require a high energy phosphate donor such as ATP or PRPP.
- Thymidine can be phosphorylated by thymidine kinase (TK).
- Uridine can be phosphorylated by uridine kinase (UK).
- Cytidine can be phosphorylated by cytidine kinase (CK).
- Deoxycytidine can be phosphorylated by deoxycytidine kinase (DCK).
Adenosine uses the enzyme adenosine kinase, which is a very important enzyme in the cell. Attempts are being made to develop an inhibitor for the enzyme for use in cancer chemotherapy.
Enzyme regulation
[edit]This protein may use the morpheein model of allosteric regulation.[4]
Clinical significance
[edit]PNPase, together with adenosine deaminase (ADA), serves a key role in purine catabolism, referred to as the salvage pathway. Mutations in ADA lead to an accumulation of (d)ATP, which inhibits ribonucleotide reductase, leading to a deficiency in (d)CTPs and (d)TTPs, which, in turn, induces apoptosis in T-lymphocytes and B-lymphocytes, leading to severe combined immunodeficiency (SCID).[citation needed]
PNP-deficient patients will have an immunodeficiency problem. It affects only T-cells; B-cells are unaffected by the deficiency.
See also
[edit]References
[edit]- ^ Canduri F, dos Santos DM, Silva RG, Mendes MA, Basso LA, Palma MS, de Azevedo WF, Santos DS (Jan 2004). "Structures of human purine nucleoside phosphorylase complexed with inosine and ddI". Biochemical and Biophysical Research Communications. 313 (4): 907–14. doi:10.1016/j.bbrc.2003.11.179. PMID 14706628.
- ^ "Entrez Gene: NP nucleoside phosphorylase".
- ^ Kaplan USMLE Biochemistry Review
- ^ Selwood T, Jaffe EK (Mar 2012). "Dynamic dissociating homo-oligomers and the control of protein function". Archives of Biochemistry and Biophysics. 519 (2): 131–43. doi:10.1016/j.abb.2011.11.020. PMC 3298769. PMID 22182754.
Further reading
[edit]- Markert ML (1991). "Purine nucleoside phosphorylase deficiency". Immunodeficiency Reviews. 3 (1): 45–81. PMID 1931007.
- Borgers M, Verhaegen H, De Brabander M, De Cree J, De Cock W, Thoné F, Geuens G (Nov 1978). "Purine nucleoside phosphorylase in chronic lymphocytic leukemia (CLL)". Blood. 52 (5): 886–95. doi:10.1182/blood.V52.5.886.886. PMID 100152.
- Aust MR, Andrews LG, Barrett MJ, Norby-Slycord CJ, Markert ML (Oct 1992). "Molecular analysis of mutations in a patient with purine nucleoside phosphorylase deficiency". American Journal of Human Genetics. 51 (4): 763–72. PMC 1682776. PMID 1384322.
- Andrews LG, Markert ML (Apr 1992). "Exon skipping in purine nucleoside phosphorylase mRNA processing leading to severe immunodeficiency". The Journal of Biological Chemistry. 267 (11): 7834–8. doi:10.1016/S0021-9258(18)42589-6. PMID 1560016.
- Jonsson JJ, Williams SR, McIvor RS (Sep 1991). "Sequence and functional characterization of the human purine nucleoside phosphorylase promoter". Nucleic Acids Research. 19 (18): 5015–20. doi:10.1093/nar/19.18.5015. PMC 328804. PMID 1923769.
- Ealick SE, Rule SA, Carter DC, Greenhough TJ, Babu YS, Cook WJ, Habash J, Helliwell JR, Stoeckler JD, Parks RE (Jan 1990). "Three-dimensional structure of human erythrocytic purine nucleoside phosphorylase at 3.2 A resolution". The Journal of Biological Chemistry. 265 (3): 1812–20. doi:10.2210/pdb2pnp/pdb. PMID 2104852.
- Williams SR, Gekeler V, McIvor RS, Martin DW (Feb 1987). "A human purine nucleoside phosphorylase deficiency caused by a single base change". The Journal of Biological Chemistry. 262 (5): 2332–8. doi:10.1016/S0021-9258(18)61658-8. PMID 3029074.
- Williams SR, Goddard JM, Martin DW (Jul 1984). "Human purine nucleoside phosphorylase cDNA sequence and genomic clone characterization". Nucleic Acids Research. 12 (14): 5779–87. doi:10.1093/nar/12.14.5779. PMC 320030. PMID 6087295.
- Pannicke U, Tuchschmid P, Friedrich W, Bartram CR, Schwarz K (Dec 1996). "Two novel missense and frameshift mutations in exons 5 and 6 of the purine nucleoside phosphorylase (PNP) gene in a severe combined immunodeficiency (SCID) patient". Human Genetics. 98 (6): 706–9. doi:10.1007/s004390050290. PMID 8931706. S2CID 5657916.
- Markert ML, Finkel BD, McLaughlin TM, Watson TJ, Collard HR, McMahon CP, Andrews LG, Barrett MJ, Ward FE (1997). "Mutations in purine nucleoside phosphorylase deficiency". Human Mutation. 9 (2): 118–21. doi:10.1002/(SICI)1098-1004(1997)9:2<118::AID-HUMU3>3.0.CO;2-5. PMID 9067751. S2CID 46243170.
- Erion MD, Takabayashi K, Smith HB, Kessi J, Wagner S, Hönger S, Shames SL, Ealick SE (Sep 1997). "Purine nucleoside phosphorylase. 1. Structure-function studies". Biochemistry. 36 (39): 11725–34. doi:10.1021/bi961969w. PMID 9305962.
- Erion MD, Stoeckler JD, Guida WC, Walter RL, Ealick SE (Sep 1997). "Purine nucleoside phosphorylase. 2. Catalytic mechanism". Biochemistry. 36 (39): 11735–48. doi:10.1021/bi961970v. PMID 9305963.
- Stoeckler JD, Poirot AF, Smith RM, Parks RE, Ealick SE, Takabayashi K, Erion MD (Sep 1997). "Purine nucleoside phosphorylase. 3. Reversal of purine base specificity by site-directed mutagenesis". Biochemistry. 36 (39): 11749–56. doi:10.1021/bi961971n. PMID 9305964.
- Sasaki Y, Iseki M, Yamaguchi S, Kurosawa Y, Yamamoto T, Moriwaki Y, Kenri T, Sasaki T, Yamashita R (Jul 1998). "Direct evidence of autosomal recessive inheritance of Arg24 to termination codon in purine nucleoside phosphorylase gene in a family with a severe combined immunodeficiency patient". Human Genetics. 103 (1): 81–5. doi:10.1007/s004390050787. PMID 9737781. S2CID 8373698.
- Sheppard TL, Ordoukhanian P, Joyce GF (Jul 2000). "A DNA enzyme with N-glycosylase activity". Proceedings of the National Academy of Sciences of the United States of America. 97 (14): 7802–7. Bibcode:2000PNAS...97.7802S. doi:10.1073/pnas.97.14.7802. PMC 16625. PMID 10884411.
- Dalal I, Grunebaum E, Cohen A, Roifman CM (Jun 2001). "Two novel mutations in a purine nucleoside phosphorylase (PNP)-deficient patient". Clinical Genetics. 59 (6): 430–7. doi:10.1034/j.1399-0004.2001.590608.x. PMID 11453975. S2CID 24624559.
- Ivings L, Pennington SR, Jenkins R, Weiss JL, Burgoyne RD (May 2002). "Identification of Ca2+-dependent binding partners for the neuronal calcium sensor protein neurocalcin delta: interaction with actin, clathrin and tubulin". The Biochemical Journal. 363 (Pt 3): 599–608. doi:10.1042/0264-6021:3630599. PMC 1222513. PMID 11964161.
- Falkenberg M, Gaspari M, Rantanen A, Trifunovic A, Larsson NG, Gustafsson CM (Jul 2002). "Mitochondrial transcription factors B1 and B2 activate transcription of human mtDNA". Nature Genetics. 31 (3): 289–94. doi:10.1038/ng909. PMID 12068295. S2CID 11164308.
- Stoychev G, Kierdaszuk B, Shugar D (Aug 2002). "Xanthosine and xanthine. Substrate properties with purine nucleoside phosphorylases, and relevance to other enzyme systems". European Journal of Biochemistry. 269 (16): 4048–57. doi:10.1046/j.1432-1033.2002.03097.x. PMID 12180982.
- Agarwal RP, Parks RE (1969). "Purine nucleoside phosphorylase from human erythrocytes. IV Crystallization and some properties". J. Biol. Chem. 244 (4): 644–7. doi:10.1016/S0021-9258(18)94403-0. PMID 5768862.
- Boyer, P.D., Lardy, H. and Myrback, K. (Eds.), The Enzymes, 2nd ed., vol. 5, Academic Press, New York, 1961, p. 237-255.
- HEPPEL LA, HILMOE RJ (1952). "[Phosphorolysis and hydrolysis of purine ribosides by enzymes from yeast.]". J. Biol. Chem. 198 (2): 683–94. doi:10.1016/S0021-9258(18)55525-3. PMID 12999785.
- Kalckar HM (1947). "The enzymatic synthesis of purine ribosides". J. Biol. Chem. 167 (2): 477–486. doi:10.1016/S0021-9258(17)31000-1. PMID 20285042.
- Saunders PP, Wilson BA, Saunders GF (1969). "Purification and comparative properties of a pyrimidine nucleoside phosphorylase from Bacillus stearothermophilus". J. Biol. Chem. 244 (13): 3691–7. doi:10.1016/S0021-9258(18)83424-X. PMID 4978445.
- Tsuboi KK, Hudson PB (1957). "Enzymes of the human erythrocyte. I. Purine nucleoside phosphorylase; isolation procedure". J. Biol. Chem. 224 (2): 879–887. doi:10.1016/S0021-9258(18)64980-4. PMID 13405917.
External links
[edit]- Human PNP at Cornell University
- E. Coli PNP at Cornell University
- Purine-Nucleoside+Phosphorylase at the U.S. National Library of Medicine Medical Subject Headings (MeSH)