HNF1A
HNF1 homeobox A (hepatocyte nuclear factor 1 homeobox A), also known as HNF1A, is a human gene on chromosome 12.[5][6][7] It is ubiquitously expressed in many tissues and cell types.[8] The protein encoded by this gene is a transcription factor that is highly expressed in the liver and is involved in the regulation of the expression of several liver-specific genes.[9] Mutations in the HNF1A gene have been known to cause diabetes.[10] The HNF1A gene also contains a SNP associated with increased risk of coronary artery disease.[11]
Structure[edit]
Gene[edit]
The HNF1A gene resides on chromosome 12 at the band 12q24.2 and contains 10[12] exons.[7] This gene produces 8 isoforms through alternative splicing.[13]
Protein[edit]
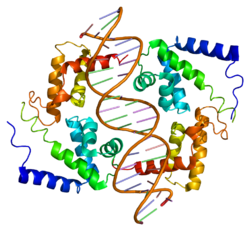
This protein belongs to the HNF1 homeobox family.[13] It contains 3 functional domains: an N-terminal dimerization domain (residues 1–32), a bipartite DNA-binding motif containing an atypical POU-homeodomain (residues 98–280), and a C-terminal transactivation domain (residues 281–631).[14][15] There is also a flexible linker (residues 33–97) which connects the dimerization and DNA binding domains.[15] Crystal structures have been solved for the dimerization domain, which forms a four-helix bundle where two α helices are separated by a turn; the DNA-binding motif, which forms a helix-turn-helix structure; and the POU-homeodomain, which is composed of three α helices, contained in the motif. This homeodomain is considered atypical due to an extended loop inserted between the second and third helices relative to the canonical homeodomain fold. The atypical insertion is thought to stabilize the interface to improve transcriptional efficiency.[14] Meanwhile, the dimerization domain is responsible for the homo- and heterodimerization of HNF-1α. The resulting dimer contains a rigid “mini-zipper”, comprising α-helices 1 and 1′, linked by a non-canonical tight turn to a flexible C-terminal comprising α-helices 2 and 2′.[15]
Function[edit]
HNF-1α is a transcription factor expressed in organs of endoderm origin, including liver, kidneys, pancreas, intestines, stomach, spleen, thymus, testis, and keratinocytes and melanocytes in human skin.[16] It has been shown to affect intestinal epithelial cell growth and cell lineages differentiation. For instance, HNF1A is an important cell-intrinsic transcription factor in adult B lymphopoiesis.[17][18][19] The participation of HNF-1α in glucose metabolism and diabetes has been reported, including the involvement in GLUT1 and GLUT2 transporter expression in pancreatic β-cells and angiotensin-converting enzyme 2 gene expression in pancreatic islets.[20][21] HNF-1α could promote the transcription of several proteins involved in the management of type II diabetes including dipeptidyl peptidase-IV (DPP-IV/CD26).[22][23] HNF-1α is also involved in various metabolic pathways of other organs, such as being a transcriptional regulator of bile acid transporters in the intestine and kidneys.[24] HNF-1α is involved in the promotion of hepatic organic cation transporters, which uptake certain classes of pharmaceuticals; hence, the loss of its function can lead to drug metabolism problems.[25] In addition, HNF-1α regulates the expression of acute phase proteins, such as fibrinogen, c-reactive protein, and interleukin 1 receptor, which are involved with inflammation.[26] Moreover, significantly lower levels of HNF-1α in pancreatic tumors and hepatocellular adenomas than in normal adjacent tissues was observed, suggesting that HNF-1α might play a possible tumor suppressor role.[27][28]
Clinical significance[edit]
HNF1A mutations can cause maturity onset diabetes of the young type 3, one of the forms of "monogenic diabetes",[6] as well as hepatocellular adenoma. HNF-1 protein is present in ovarian clear-cell carcinoma.[29][30]
In humans, mutations in HNF1A cause diabetes that responds to low dose sulfonylurea agents.[31] The identification of extreme sulfonylurea sensitivity in patients with diabetes mellitus owing to heterozygous mutations in HNF1A presents a clear example of the relevance of HNF1A in diabetes patients and how pharmacogenetics can contribute in patient care.[32] For example, patients with maturity onset diabetes of the young owing to mutations in HNF1A (which accounts for ~3% of all diabetes mellitus cases diagnosed under the age of 30 years) are extremely sensitive to sulfonylurea treatment and can successfully transition off insulin treatment.[10] Likewise, patients with diabetes caused by mutations in the HNF1A gene have been described as sensitive to the hypoglycemic effects of sulphonylureas. The cause of hyperglycemia appears to alter the response to hypoglycemic drugs. Accordingly, HNF-1α-induced diabetes has marked sulphonylurea sensitivity. This pharmacogenetic effect is consistent with models of HNF-1α deficiency, and the genetic basis of hyperglycemia may have implications for patient management.[10] Common genetic variation within HNF1A is also associated with risk of developing type 2 diabetes and increased penetrance of early-onset diabetes[33]
Clinical marker[edit]
A multi-locus genetic risk score study based on a combination of 27 loci, including the HNF1A gene, identified individuals at increased risk for both incident and recurrent coronary artery disease events, as well as an enhanced clinical benefit from statin therapy. The study was based on a community cohort study (the Malmo Diet and Cancer study) and four additional randomized controlled trials of primary prevention cohorts (JUPITER and ASCOT) and secondary prevention cohorts (CARE and PROVE IT-TIMI 22).[11]
Interactions[edit]
HNF1A has been shown to interact with:
See also[edit]
- Hepatocyte nuclear factors
- Maturity onset diabetes of the young
- Other HNF class homeoboxes:
References[edit]
- ^ a b c GRCh38: Ensembl release 89: ENSG00000135100 – Ensembl, May 2017
- ^ a b c GRCm38: Ensembl release 89: ENSMUSG00000029556 – Ensembl, May 2017
- ^ "Human PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- ^ "Mouse PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- ^ Szpirer C, Riviere M, Cortese R, Nakamura T, Islam MQ, Levan G, Szpirer J (June 1992). "Chromosomal localization in man and rat of the genes encoding the liver-enriched transcription factors C/EBP, DBP, and HNF1/LFB-1 (CEBP, DBP, and transcription factor 1, TCF1, respectively) and of the hepatocyte growth factor/scatter factor gene (HGF)". Genomics. 13 (2): 293–300. doi:10.1016/0888-7543(92)90245-N. PMID 1535333.
- ^ a b Vaxillaire M, Boccio V, Philippi A, Vigouroux C, Terwilliger J, Passa P, Beckmann JS, Velho G, Lathrop GM, Froguel P (April 1995). "A gene for maturity onset diabetes of the young (MODY) maps to chromosome 12q". Nature Genetics. 9 (4): 418–23. doi:10.1038/ng0495-418. PMID 7795649. S2CID 665243.
- ^ a b "HNF1A HNF1 homeobox A [Homo sapiens (human)] - Gene - NCBI". www.ncbi.nlm.nih.gov. Retrieved 2016-10-11.
- ^ "BioGPS - your Gene Portal System". biogps.org. Retrieved 2016-10-11.
- ^ Courtois G, Morgan JG, Campbell LA, Fourel G, Crabtree GR (October 1987). "Interaction of a liver-specific nuclear factor with the fibrinogen and alpha 1-antitrypsin promoters". Science. 238 (4827): 688–92. Bibcode:1987Sci...238..688C. doi:10.1126/science.3499668. PMID 3499668.
- ^ a b c Pearson ER, Starkey BJ, Powell RJ, Gribble FM, Clark PM, Hattersley AT (October 2003). "Genetic cause of hyperglycaemia and response to treatment in diabetes". Lancet. 362 (9392): 1275–81. doi:10.1016/S0140-6736(03)14571-0. PMID 14575972. S2CID 34914098.
- ^ a b Mega JL, Stitziel NO, Smith JG, Chasman DI, Caulfield MJ, Devlin JJ, Nordio F, Hyde CL, Cannon CP, Sacks FM, Poulter NR, Sever PS, Ridker PM, Braunwald E, Melander O, Kathiresan S, Sabatine MS (June 2015). "Genetic risk, coronary heart disease events, and the clinical benefit of statin therapy: an analysis of primary and secondary prevention trials". Lancet. 385 (9984): 2264–71. doi:10.1016/S0140-6736(14)61730-X. PMC 4608367. PMID 25748612.
- ^ "Homo sapiens HNF1 homeobox A (HNF1A), RefSeqGene (LRG_522) on chromosome 12". 13 June 2021.
- ^ a b "HNF1A - Hepatocyte nuclear factor 1-alpha - Homo sapiens (Human) - HNF1A gene & protein". www.uniprot.org. Retrieved 2016-10-11.
- ^ a b Chi YI, Frantz JD, Oh BC, Hansen L, Dhe-Paganon S, Shoelson SE (November 2002). "Diabetes mutations delineate an atypical POU domain in HNF-1alpha". Molecular Cell. 10 (5): 1129–37. doi:10.1016/s1097-2765(02)00704-9. PMID 12453420.
- ^ a b c Narayana N, Phillips NB, Hua QX, Jia W, Weiss MA (September 2006). "Diabetes mellitus due to misfolding of a beta-cell transcription factor: stereospecific frustration of a Schellman motif in HNF-1alpha". Journal of Molecular Biology. 362 (3): 414–29. doi:10.1016/j.jmb.2006.06.086. PMID 16930618.
- ^ Real Hernandez LM, Fan J, Johnson MH, Gonzalez de Mejia E (2015-01-01). "Berry Phenolic Compounds Increase Expression of Hepatocyte Nuclear Factor-1α (HNF-1α) in Caco-2 and Normal Colon Cells Due to High Affinities with Transcription and Dimerization Domains of HNF-1α". PLOS ONE. 10 (9): e0138768. Bibcode:2015PLoSO..1038768R. doi:10.1371/journal.pone.0138768. PMC 4587667. PMID 26413797.
- ^ Lussier CR, Brial F, Roy SA, Langlois MJ, Verdu EF, Rivard N, Perreault N, Boudreau F (2010-01-01). "Loss of hepatocyte-nuclear-factor-1alpha impacts on adult mouse intestinal epithelial cell growth and cell lineages differentiation". PLOS ONE. 5 (8): e12378. Bibcode:2010PLoSO...512378L. doi:10.1371/journal.pone.0012378. PMC 2927538. PMID 20808783.
- ^ D'Angelo A, Bluteau O, Garcia-Gonzalez MA, Gresh L, Doyen A, Garbay S, Robine S, Pontoglio M (May 2010). "Hepatocyte nuclear factor 1alpha and beta control terminal differentiation and cell fate commitment in the gut epithelium". Development. 137 (9): 1573–82. doi:10.1242/dev.044420. PMID 20388655.
- ^ von Wnuck Lipinski K, Sattler K, Peters S, Weske S, Keul P, Klump H, Heusch G, Göthert JR, Levkau B (February 2016). "Hepatocyte Nuclear Factor 1A Is a Cell-Intrinsic Transcription Factor Required for B Cell Differentiation and Development in Mice". Journal of Immunology. 196 (4): 1655–65. doi:10.4049/jimmunol.1500897. PMID 26800876.
- ^ Luni C, Marth JD, Doyle FJ (2012-01-01). "Computational modeling of glucose transport in pancreatic β-cells identifies metabolic thresholds and therapeutic targets in diabetes". PLOS ONE. 7 (12): e53130. Bibcode:2012PLoSO...753130L. doi:10.1371/journal.pone.0053130. PMC 3531366. PMID 23300881.
- ^ Pedersen KB, Chhabra KH, Nguyen VK, Xia H, Lazartigues E (November 2013). "The transcription factor HNF1α induces expression of angiotensin-converting enzyme 2 (ACE2) in pancreatic islets from evolutionarily conserved promoter motifs". Biochimica et Biophysica Acta (BBA) - Gene Regulatory Mechanisms. 1829 (11): 1225–35. doi:10.1016/j.bbagrm.2013.09.007. PMC 3838857. PMID 24100303.
- ^ Gu N, Adachi T, Matsunaga T, Takeda J, Tsujimoto G, Ishihara A, Yasuda K, Tsuda K (August 2006). "Mutant HNF-1alpha and mutant HNF-1beta identified in MODY3 and MODY5 downregulate DPP-IV gene expression in Caco-2 cells". Biochemical and Biophysical Research Communications. 346 (3): 1016–23. doi:10.1016/j.bbrc.2006.06.010. PMID 16781669.
- ^ Gu N, Tsuda M, Matsunaga T, Adachi T, Yasuda K, Ishihara A, Tsuda K (December 2008). "Glucose regulation of dipeptidyl peptidase IV gene expression is mediated by hepatocyte nuclear factor-1alpha in epithelial intestinal cells". Clinical and Experimental Pharmacology & Physiology. 35 (12): 1433–9. doi:10.1111/j.1440-1681.2008.05015.x. PMID 18671716. S2CID 24464350.
- ^ Shih DQ, Bussen M, Sehayek E, Ananthanarayanan M, Shneider BL, Suchy FJ, Shefer S, Bollileni JS, Gonzalez FJ, Breslow JL, Stoffel M (April 2001). "Hepatocyte nuclear factor-1alpha is an essential regulator of bile acid and plasma cholesterol metabolism". Nature Genetics. 27 (4): 375–82. doi:10.1038/86871. PMID 11279518. S2CID 22640762.
- ^ O'Brien VP, Bokelmann K, Ramírez J, Jobst K, Ratain MJ, Brockmöller J, Tzvetkov MV (October 2013). "Hepatocyte nuclear factor 1 regulates the expression of the organic cation transporter 1 via binding to an evolutionary conserved region in intron 1 of the OCT1 gene". The Journal of Pharmacology and Experimental Therapeutics. 347 (1): 181–92. doi:10.1124/jpet.113.206359. PMC 3781413. PMID 23922447.
- ^ Armendariz AD, Krauss RM (April 2009). "Hepatic nuclear factor 1-alpha: inflammation, genetics, and atherosclerosis". Current Opinion in Lipidology. 20 (2): 106–11. doi:10.1097/mol.0b013e3283295ee9. PMID 19280766. S2CID 85192269.
- ^ Luo Z, Li Y, Wang H, Fleming J, Li M, Kang Y, Zhang R, Li D (2015-01-01). "Hepatocyte nuclear factor 1A (HNF1A) as a possible tumor suppressor in pancreatic cancer". PLOS ONE. 10 (3): e0121082. Bibcode:2015PLoSO..1021082L. doi:10.1371/journal.pone.0121082. PMC 4368635. PMID 25793983.
- ^ Bluteau O, Jeannot E, Bioulac-Sage P, Marqués JM, Blanc JF, Bui H, Beaudoin JC, Franco D, Balabaud C, Laurent-Puig P, Zucman-Rossi J (October 2002). "Bi-allelic inactivation of TCF1 in hepatic adenomas". Nature Genetics. 32 (2): 312–5. doi:10.1038/ng1001. PMID 12355088. S2CID 11919671.
- ^ Köbel M, Kalloger SE, Carrick J, Huntsman D, Asad H, Oliva E, Ewanowich CA, Soslow RA, Gilks CB (January 2009). "A limited panel of immunomarkers can reliably distinguish between clear cell and high-grade serous carcinoma of the ovary". The American Journal of Surgical Pathology. 33 (1): 14–21. doi:10.1097/PAS.0b013e3181788546. PMID 18830127. S2CID 19610205.
- ^ Offman SL, Longacre TA (September 2012). "Clear cell carcinoma of the female genital tract (not everything is as clear as it seems)". Advances in Anatomic Pathology. 19 (5): 296–312. doi:10.1097/PAP.0b013e31826663b1. PMID 22885379. S2CID 33159622.
- ^ Owen KR (June 2016). "Treating young adults with type 2 diabetes or monogenic diabetes". Best Practice & Research. Clinical Endocrinology & Metabolism. 30 (3): 455–67. doi:10.1016/j.beem.2016.05.002. PMID 27432078.
- ^ Zhou K, Pedersen HK, Dawed AY, Pearson ER (June 2016). "Pharmacogenomics in diabetes mellitus: insights into drug action and drug discovery" (PDF). Nature Reviews. Endocrinology. 12 (6): 337–46. doi:10.1038/nrendo.2016.51. PMID 27062931. S2CID 20405132.
- ^ "The Common HNF1A Variant I27L Is a Modifier of Age at Diabetes Diagnosis in Individuals With HNF1A-MODY"
- ^ a b c d Soutoglou E, Papafotiou G, Katrakili N, Talianidis I (April 2000). "Transcriptional activation by hepatocyte nuclear factor-1 requires synergism between multiple coactivator proteins". The Journal of Biological Chemistry. 275 (17): 12515–20. doi:10.1074/jbc.275.17.12515. PMID 10777539.
- ^ Ban N, Yamada Y, Someya Y, Miyawaki K, Ihara Y, Hosokawa M, Toyokuni S, Tsuda K, Seino Y (May 2002). "Hepatocyte nuclear factor-1alpha recruits the transcriptional co-activator p300 on the GLUT2 gene promoter". Diabetes. 51 (5): 1409–18. doi:10.2337/diabetes.51.5.1409. PMID 11978637.
- ^ Ewing RM, Chu P, Elisma F, Li H, Taylor P, Climie S, McBroom-Cerajewski L, Robinson MD, O'Connor L, Li M, Taylor R, Dharsee M, Ho Y, Heilbut A, Moore L, Zhang S, Ornatsky O, Bukhman YV, Ethier M, Sheng Y, Vasilescu J, Abu-Farha M, Lambert JP, Duewel HS, Stewart II, Kuehl B, Hogue K, Colwill K, Gladwish K, Muskat B, Kinach R, Adams SL, Moran MF, Morin GB, Topaloglou T, Figeys D (2007). "Large-scale mapping of human protein-protein interactions by mass spectrometry". Molecular Systems Biology. 3: 89. doi:10.1038/msb4100134. PMC 1847948. PMID 17353931.
- ^ Sourdive DJ, Transy C, Garbay S, Yaniv M (April 1997). "The bifunctional DCOH protein binds to HNF1 independently of its 4-alpha-carbinolamine dehydratase activity". Nucleic Acids Research. 25 (8): 1476–84. doi:10.1093/nar/25.8.1476. PMC 146627. PMID 9092652.
- ^ Ficner, R.; Sauer, U.H.; Stier, G.; Suck, D. (May 1995). "Three-dimensional structure of the bifunctional protein PCD/DCoH, a cytoplasmic enzyme interacting with transcription factor HNF1". The EMBO Journal. 14 (9): 2034–2042. doi:10.1002/j.1460-2075.1995.tb07195.x. hdl:11858/00-001M-0000-0019-A7D6-3. PMC 398303. PMID 7744010.
- ^ Alber, Tom; Rose, Robert B.; Bayle, J. Henri; Endrizzi, James A.; Cronk, Jeff D.; Crabtree, Gerald R. (2000-09-01). "Structural basis of dimerization, coactivator recognition and MODY3 mutations in HNF-1alpha". Nature Structural Biology. 7 (9): 744–748. doi:10.1038/78966. PMID 10966642. S2CID 22528806.
- ^ Hines RN, Luo Z, Cresteil T, Ding X, Prough RA, Fitzpatrick JL, Ripp SL, Falkner KC, Ge NL, Levine A, Elferink CJ (May 2001). "Molecular regulation of genes encoding xenobiotic-metabolizing enzymes: mechanisms involving endogenous factors". Drug Metabolism and Disposition. 29 (5): 623–33. PMID 11302926.
- ^ Rufibach LE, Duncan SA, Battle M, Deeb SS (July 2006). "Transcriptional regulation of the human hepatic lipase (LIPC) gene promoter". Journal of Lipid Research. 47 (7): 1463–77. doi:10.1194/jlr.M600082-JLR200. PMID 16603721.
- ^ a b Bulla GA, Kraus DM (December 2004). "Dissociation of the hepatic phenotype from HNF4 and HNF1alpha expression". Bioscience Reports. 24 (6): 595–608. doi:10.1007/s10540-005-2794-8. PMID 16158197. S2CID 55553843.
Further reading[edit]
- Eastman Q, Grosschedl R (April 1999). "Regulation of LEF-1/TCF transcription factors by Wnt and other signals". Current Opinion in Cell Biology. 11 (2): 233–40. doi:10.1016/S0955-0674(99)80031-3. PMID 10209158.
- Bioulac-Sage P, Blanc JF, Rebouissou S, Balabaud C, Zucman-Rossi J (May 2007). "Genotype phenotype classification of hepatocellular adenoma". World Journal of Gastroenterology. 13 (19): 2649–54. doi:10.3748/wjg.v13.i19.2649. PMC 4147112. PMID 17569132.