RUNX1
Runt-related transcription factor 1 (RUNX1) also known as acute myeloid leukemia 1 protein (AML1) or core-binding factor subunit alpha-2 (CBFA2) is a protein that in humans is encoded by the RUNX1 gene.[5][6]
RUNX1 is a transcription factor that regulates the differentiation of hematopoietic stem cells into mature blood cells.[7] In addition it plays a major role in the development of the neurons that transmit pain.[8] It belongs to the Runt-related transcription factor (RUNX) family of genes which are also called core binding factor-α (CBFα). RUNX proteins form a heterodimeric complex with CBFβ which confers increased DNA binding and stability to the complex.
Chromosomal translocations involving the RUNX1 gene are associated with several types of leukemia including M2 AML.[9] Mutations in RUNX1 are implicated in cases of breast cancer.[10]
Gene and protein[edit]
In humans, the gene RUNX1 is 260 kilobases (kb) in length, and is located on chromosome 21 (21q22.12). The gene can be transcribed from 2 alternative promoters, promoter 1 (distal) or promoter 2 (proximal). As a result, various isoforms of RUNX1 can be synthesized, facilitated by alternative splicing. The full-length RUNX1 protein is encoded by 12 exons. Among the exons are two defined domains, namely the runt homology domain (RHD) or the runt domain (exons 2, 3 and 4), and the transactivation domain (TAD) (exon 6). These domains are necessary for RUNX1 to mediate DNA binding and protein-protein interactions respectively. The transcription of RUNX1 is regulated by 2 enhancers (regulatory element 1 and regulatory element 2), and these tissue specific enhancers enable the binding of lymphoid or erythroid regulatory proteins, therefore the gene activity of RUNX1 is highly active in the haematopoietic system.
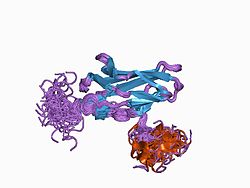
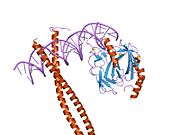
The protein RUNX1 is composed of 453 amino acids. As a transcription factor (TF), its DNA binding ability is encoded by the runt domain (residues 50 – 177), which is homologous to the p53 family. The runt domain of RUNX1 binds to the core consensus sequence TGTGGNNN (where NNN can represent either TTT or TCA).[11] DNA recognition is achieved by loops of the 12-stranded β-barrel and the C-terminus “tail” (residues 170 – 177), which clamp around the sugar phosphate backbone and fits into the major and minor grooves of DNA. Specificity is achieved by making direct or water-mediated contacts with the bases. RUNX1 can bind DNA as a monomer, but its DNA binding affinity is enhanced by 10 fold if it heterodimerises with the core binding factor β (CBFβ), also via the runt domain. In fact, the RUNX family is often referred to as α-subunits, together with binding of a common β-subunit CBFβ, RUNX can behave as heterodimeric transcription factors collectively called the core binding factors (CBFs).
The consensus binding site for CBF has been identified to be a 7 bp sequence PyGPyGGTPy. Py denotes pyrimidine which can be either cytosine or thymine.[12]
Discovery and characterization of RUNX1[edit]
Nusslein-Volhard and Wieschaus discovered the transcription factor RUNX in a screen that was conducted to identify mutations that affect segment number and polarity in Drosophila.[13] The mutation that led to presegmentation patterning defects and runted embryos was named runt. Following this discovery, the Drosophila segmentation gene runt was cloned by Gergen et al. Although the protein encoded by runt was demonstrated to exhibit nuclear translocation, it was not yet established that this protein is a transcription factor.[14] Subsequently, in 1991, Ohki et al. cloned the human RUNX1 gene; RUNX1 was found to be rearranged in the leukemic cell DNAs from t(8;21)(q22;q22) AML patients.[15] However, the function of human RUNX1 was not established. Soon after the discovery of the drosophila runt protein and the human RUNX1 protein, RUNX1's function was discovered. Runx1 was purified as a sequence-specific DNA-binding protein that regulated the disease specificity of the Moloney murine Leukemia virus.[16] Furthermore, Ito et al. purified Runx2, the homolog of Runx1.[17] Purified transcription factors consisted of two subunits, a DNA binding CBFα chain (RUNX1 or RUNX2) and a non-DNA-binding subunit called core binding factor β (CBFβ); the binding affinity of RUNX1 and RUNX2 was significantly increased by association with CBFβ.[17][18][19]
Mouse knockout[edit]
Mice embryos with homozygous mutations on RUNX1 died at about 12.5 days. The embryos displayed lack of fetal liver hematopoiesis.[20]
Similar experiments from a different research group demonstrated that the knockout embryos die between embryonic days 11.5 and 12.5 due to hemorrhaging in the central nervous system (CNS).[21]
Participation in haematopoiesis[edit]
RUNX1 plays a crucial role in adult (definitive) haematopoiesis during embryonic development. It is expressed in all haematopoietic sites that contribute to the formation of haematopoietic stem and progenitor cells (HSPCs), including the yolk sac,[22] allantois, placenta, para-aortic splanchnopleura (P-Sp; (the visceral mesodermal layer),[23] aorta-gonad-mesonephros (AGM) and the umbilical and vitelline arteries.[24] HSPCs are generated via the hemogenic endothelium, a special subset of endothelial cells scattered within blood vessels that can differentiate into haematopoietic cells. The emergence of HSPCs is often studied in mouse and zebrafish animal models, in which HSPCs appear as “intra-aortic” clusters that adhere to the ventral wall of the dorsal aorta. RUNX1 or CBF takes part in this process by mediating the transition of an endothelial cell to become a haematopoietic cell.[25] There is increasing evidence that RUNX1 may also be important during primitive haematopoiesis.[26] This is because in RUNX1 knockout mice, primitive erythrocytes displayed a defective morphology and the size of blast cell population was substantially reduced, apart from the absence of HSPCs which would result in embryonic lethality by Embryonic day (E) 11.5 – 12.5.
At a molecular level, expression of the gene RUNX1 is upregulated by the RUNX1 intronic cis-regulatory element (+23 RUNX1 enhancer).[27] This +23 RUNX1 enhancer contains conserved motifs that encourage binding of various haematopoiesis related regulators such as Gata2, ETS factors (Fli-1, Elf-1, PU.1) and the SCL / Lmo2 / Ldb1 complex, as well as RUNX1 itself acting in an auto-regulatory loop. As mentioned before, the main role of RUNX1 is to modulate the fate of haematopoietic cells. This can be achieved by binding to the thrombopoietin (TPO) receptor/ c-Mpl promoter, followed by the recruitment of transcription activators or repressors in order to promote transition of the hemogenic endothelium to HSCs, or differentiation into lineages of lower haematopoietic hierarchies. RUNX1 can also modulate its own level by upregulating the expression of Smad6 to target itself for proteolysis.[28]
Mutations and acute myeloid leukemia[edit]
A broad range of heterozygous germline mutations in RUNX1 have been associated with Familial Platelet Disorder, a mild bleeding disorder associated with a high rate of myeloid leukemia.[29] At least 39 forms of somatic RUNX1 mutation are implicated in various myeloid malignancies. Examples range from RUNX1 point mutations acquired from low-dose radiation leading to myelodysplastic neoplasms or therapy-related myeloid neoplasms, to chromosomal translocation of the RUNX1 gene with the ETO / MTG8 / RUNX1T1 gene located on chromosome 8q22, t(8; 21), generating a fusion protein AML-ETO, categorized as acute myeloid leukemia (AML) M2.
In t(8; 21), breakpoints frequently occur at intron 5 – 6 of RUNX1 and intron 1b – 2 of ETO, creating chimeric transcripts that inherit the runt domain from RUNX1, and all Nervy homology regions (NHR) 1-4 from ETO. As a consequence, AML-ETO retains the ability to bind at RUNX1 target genes whilst acting as a transcription repressor via the recruitment of corepressors and histone deacetylases, which is an intrinsic function of ETO. Oncogenic potential of AML-ETO is exerted because it blocks differentiation and promote self-renewal in blast cells, resulting in massive accumulation of blasts (>20%) in the bone marrow. This is further characterized histologically by the presence of Auer rods and epigenetically by lysine acetylation on residues 24 and 43. Other actions of AML-ETO that could induce leukemogenesis include downregulation of the DNA repair enzyme 8-oxoguanine DNA glycosylase (OGG1) and increase in the level of intracellular reactive oxygen species, making cells that express AML-ETO more susceptible to additional genetic mutations.
Role in T-cell acute lymphoblastic leukemia (T-ALL)[edit]
Around 15% of T-ALL patients have RUNX1 mutations which are clustered around the DNA binding domain of RUNX1. Those mutations are proposed to cause loss-of-function and might play a tumor suppressor role.[30]
Participation in hair follicle development[edit]
Runx1 was first discovered to be expressed in mouse embryonic skin.[31] It is expressed in the epithelial compartment to control hair follicle activation from telogen to anagen through activating Wnt signaling and Lef1 levels [32] At the same time it is expressed in the dermis where it suppresses the same targets to allow for embryogenic development of hair shaft and follicles.[33] In the human hair follicle the expression patterns are similar to the mouse - indicating that it plays a similar role.[34] In addition to hair follicle development, Runx1 is also implicated in skin and epithelial cancer development.[34][35] Thus there are similarities across tissue in Runx1 behavior.
RUNX1 in Pancreatic Cancer[edit]
High expression of RUNX1 is associated with adverse survival of pancreatic cancer patients and has tumor promoting potential in pancreatic cancer.[36] The most common cause of resistance to therapeutic treatments is the suppression of the programmed cell death (apoptosis) of pancreatic cancer cells. A key factor in apoptosis initiation is the protein NOXA, which is suppressed in a particularly aggressive form of pancreatic cancer. Genetic suppression of the NOXA gene is mediated by the transcription factor RUNX1. Pharmacological or genetic inhibition of RUNX1 de-represses the NOXA gene and induces apoptosis in pancreatic cancer cells.[36]
Interactions[edit]
RUNX1 has been shown to interact with:
See also[edit]
References[edit]
- ^ a b c GRCh38: Ensembl release 89: ENSG00000159216 – Ensembl, May 2017
- ^ a b c GRCm38: Ensembl release 89: ENSMUSG00000022952 – Ensembl, May 2017
- ^ "Human PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- ^ "Mouse PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- ^ "Entrez Gene: RUNX1 runt-related transcription factor 1 (acute myeloid leukemia 1; aml1 oncogene)".
- ^ Avramopoulos D, Cox T, Blaschak JE, Chakravarti A, Antonarakis SE (October 1992). "Linkage mapping of the AML1 gene on human chromosome 21 using a DNA polymorphism in the 3' untranslated region". Genomics. 14 (2): 506–7. doi:10.1016/S0888-7543(05)80253-8. PMID 1427868.
- ^ Okuda T, Nishimura M, Nakao M, Fujita Y (October 2001). "RUNX1/AML1: a central player in hematopoiesis". International Journal of Hematology. 74 (3): 252–7. doi:10.1007/bf02982057. PMID 11721959. S2CID 5918511.
- ^ Chen CL, Broom DC, Liu Y, de Nooij JC, Li Z, Cen C, Samad OA, Jessell TM, Woolf CJ, Ma Q (February 2006). "Runx1 determines nociceptive sensory neuron phenotype and is required for thermal and neuropathic pain". Neuron. 49 (3): 365–77. doi:10.1016/j.neuron.2005.10.036. PMID 16446141. S2CID 16070223.
- ^ Asou N (February 2003). "The role of a Runt domain transcription factor AML1/RUNX1 in leukemogenesis and its clinical implications". Critical Reviews in Oncology/Hematology. 45 (2): 129–50. doi:10.1016/S1040-8428(02)00003-3. PMID 12604126.
- ^ Koboldt DC (October 2012). "Comprehensive molecular portraits of human breast tumours". Nature. 490 (7418). Nature Publishing Group: 61–70. Bibcode:2012Natur.490...61T. doi:10.1038/nature11412. PMC 3465532. PMID 23000897.
- ^ Bowers SR, Calero-Nieto FJ, Valeaux S, Fernandez-Fuentes N, Cockerill PN (October 2010). "Runx1 binds as a dimeric complex to overlapping Runx1 sites within a palindromic element in the human GM-CSF enhancer". Nucleic Acids Research. 38 (18): 6124–34. doi:10.1093/nar/gkq356. PMC 2952845. PMID 20483917.
- ^ Melnikova IN, Crute BE, Wang S, Speck NA (April 1993). "Sequence specificity of the core-binding factor". Journal of Virology. 67 (4): 2408–11. doi:10.1128/JVI.67.4.2408-2411.1993. PMC 240414. PMID 8445737.
- ^ Nüsslein-Volhard, C, Wieschaus, E (October 1980). "Mutations affecting segment number and polarity in Drosophila". Nature. 287 (5785): 795–801. Bibcode:1980Natur.287..795N. doi:10.1038/287795a0. PMID 6776413. S2CID 4337658.
- ^ Kania, MA, Bonner, AS, Duffy, JB, Gergen, JP (October 1990). "The Drosophila segmentation gene runt encodes a novel nuclear regulatory protein that is also expressed in the developing nervous system". Genes Dev. 4 (10): 1701–1713. doi:10.1101/gad.4.10.1701. PMID 2249771.
- ^ Miyoshi, H, Shimizu, K, Kozu, T, Maseki, N, Kaneko, Y, Ohki, M (December 1991). "t(8;21) breakpoints on chromosome 21 in acute myeloid leukemia are clustered within a limited region of a single gene, AML1". Proc Natl Acad Sci USA. 88 (23): 10431–10434. Bibcode:1991PNAS...8810431M. doi:10.1073/pnas.88.23.10431. PMC 52942. PMID 1720541.
- ^ Wang, S, Speck, NA (January 1992). "Purification of core-binding factor, a protein that binds the conserved core site in murine leukemia virus enhancers". Mol Cell Biol. 12 (1): 89–102. doi:10.1128/MCB.12.1.89. PMC 364072. PMID 1309596.
- ^ a b Kamachi Y, Ogawa E, Asano M, Ishida S, Murakami Y, Satake M, Ito Y, Shigesada K (October 1990). "Purification of a mouse nuclear factor that binds to both the A and B cores of the polyomavirus enhancer". J Virol. 64 (10): 4808–4819. doi:10.1128/JVI.64.10.4808-4819.1990. PMC 247969. PMID 2168969.
- ^ Ogawa E, Inuzuka M, Maruyama M, Satake M, Naito-Fujimoto M, Ito Y, Shigesada K (May 1993). "Molecular cloning and characterization of PEBP2 beta, the heterodimeric partner of a novel Drosophila runt-related DNA binding protein PEBP2 alpha". Virology. 194 (1): 314–331. doi:10.1006/viro.1993.1262. PMID 8386878.
- ^ Wang, S, Wang, Q, Crute, BE, Melnikova, IN, Keller, SR, Speck, NA (June 1993). "Cloning and characterization of subunits of the T-cell receptor and murine leukemia virus enhancer core-binding factor". Mol Cell Biol. 13 (6): 3324–39. doi:10.1128/MCB.13.6.3324. PMC 359789. PMID 8497254.
- ^ Okuda T, van Deursen J, Hiebert SW, Grosveld G, Downing JR (January 1996). "AML1, the target of multiple chromosomal translocations in human leukemia, is essential for normal fetal liver hematopoiesis". Cell. 84 (2): 321–30. doi:10.1016/S0092-8674(00)80986-1. PMID 8565077. S2CID 14180316.
- ^ Wang Q, Stacy T, Binder M, Marin-Padilla M, Sharpe AH, Speck NA (April 1996). "Disruption of the Cbfa2 gene causes necrosis and hemorrhaging in the central nervous system and blocks definitive hematopoiesis". Proceedings of the National Academy of Sciences of the United States of America. 93 (8): 3444–9. Bibcode:1996PNAS...93.3444W. doi:10.1073/pnas.93.8.3444. PMC 39628. PMID 8622955.
- ^ Yoshimoto M, Montecino-Rodriguez E, Ferkowicz MJ, Porayette P, Shelley WC, Conway SJ, et al. (January 2011). "Embryonic day 9 yolk sac and intra-embryonic hemogenic endothelium independently generate a B-1 and marginal zone progenitor lacking B-2 potential". Proceedings of the National Academy of Sciences of the United States of America. 108 (4): 1468–1473. Bibcode:2011PNAS..108.1468Y. doi:10.1073/pnas.1015841108. PMC 3029764. PMID 21209332.
- ^ Godin IE, Garcia-Porrero JA, Coutinho A, Dieterlen-Lièvre F, Marcos MA (July 1993). "Para-aortic splanchnopleura from early mouse embryos contains B1a cell progenitors". Nature. 364 (6432): 67–70. Bibcode:1993Natur.364...67G. doi:10.1038/364067a0. PMID 8316299. S2CID 4254064.
- ^ Lin Y, Yoder MC, Yoshimoto M (June 2014). "Lymphoid progenitor emergence in the murine embryo and yolk sac precedes stem cell detection". Stem Cells and Development. 23 (11): 1168–1177. doi:10.1089/scd.2013.0536. PMC 4028089. PMID 24417306.
- ^ Chen MJ, Yokomizo T, Zeigler BM, Dzierzak E, Speck NA (February 2009). "Runx1 is required for the endothelial to haematopoietic cell transition but not thereafter". Nature. 457 (7231): 887–891. Bibcode:2009Natur.457..887C. doi:10.1038/nature07619. PMC 2744041. PMID 19129762.
- ^ Yokomizo T, Hasegawa K, Ishitobi H, Osato M, Ema M, Ito Y, et al. (April 2008). "Runx1 is involved in primitive erythropoiesis in the mouse" (PDF). Blood. 111 (8): 4075–4080. doi:10.1182/blood-2007-05-091637. PMID 18250229.
- ^ Nottingham WT, Jarratt A, Burgess M, Speck CL, Cheng JF, Prabhakar S, et al. (December 2007). "Runx1-mediated hematopoietic stem-cell emergence is controlled by a Gata/Ets/SCL-regulated enhancer". Blood. 110 (13): 4188–4197. doi:10.1182/blood-2007-07-100883. PMC 2234795. PMID 17823307.
- ^ Knezevic K, Bee T, Wilson NK, Janes ME, Kinston S, Polderdijk S, et al. (July 2011). "A Runx1-Smad6 rheostat controls Runx1 activity during embryonic hematopoiesis". Molecular and Cellular Biology. 31 (14): 2817–2826. doi:10.1128/MCB.01305-10. PMC 3133398. PMID 21576367.
- ^ Sood R, Kamikubo Y, Liu P (April 2017). "Role of RUNX1 in hematological malignancies". Blood. 129 (15): 2070–2082. doi:10.1182/blood-2016-10-687830. PMC 5391618. PMID 28179279.
- ^ Grossmann V, Kern W, Harbich S, Alpermann T, Jeromin S, Schnittger S, et al. (December 2011). "Prognostic relevance of RUNX1 mutations in T-cell acute lymphoblastic leukemia". Haematologica. 96 (12): 1874–1877. doi:10.3324/haematol.2011.043919. PMC 3232273. PMID 21828118.
- ^ North TE, de Bruijn MF, Stacy T, Talebian L, Lind E, Robin C, Binder M, Dzierzak E, Speck NA (May 2002). "Runx1 expression marks long-term repopulating hematopoietic stem cells in the midgestation mouse embryo". Immunity. 16 (5): 661–72. doi:10.1016/s1074-7613(02)00296-0. PMID 12049718.
- ^ Osorio KM, Lee SE, McDermitt DJ, Waghmare SK, Zhang YV, Woo HN, Tumbar T (March 2008). "Runx1 modulates developmental, but not injury-driven, hair follicle stem cell activation". Development. 135 (6): 1059–68. doi:10.1242/dev.012799. PMID 18256199.
- ^ Osorio KM, Lilja KC, Tumbar T (April 2011). "Runx1 modulates adult hair follicle stem cell emergence and maintenance from distinct embryonic skin compartments". The Journal of Cell Biology. 193 (1): 235–50. doi:10.1083/jcb.201006068. PMC 3082184. PMID 21464233.
- ^ a b c Scheitz CJ, Lee TS, McDermitt DJ, Tumbar T (November 2012). "Defining a tissue stem cell-driven Runx1/Stat3 signalling axis in epithelial cancer". The EMBO Journal. 31 (21): 4124–39. doi:10.1038/emboj.2012.270. PMC 3492731. PMID 23034403.
- ^ Hoi CS, Lee SE, Lu SY, McDermitt DJ, Osorio KM, Piskun CM, Peters RM, Paus R, Tumbar T (May 2010). "Runx1 directly promotes proliferation of hair follicle stem cells and epithelial tumor formation in mouse skin". Molecular and Cellular Biology. 30 (10): 2518–36. doi:10.1128/MCB.01308-09. PMC 2863705. PMID 20308320.
- ^ a b Doffo J, Bamopoulos SA, Köse H, Orben F, Zang C, Pons M, et al. (March 2022). "NOXA expression drives synthetic lethality to RUNX1 inhibition in pancreatic cancer". Proceedings of the National Academy of Sciences of the United States of America. 119 (9): e2105691119. Bibcode:2022PNAS..11905691D. doi:10.1073/pnas.2105691119. PMC 8892327. PMID 35197278.
- ^ a b Hess J, Porte D, Munz C, Angel P (June 2001). "AP-1 and Cbfa/runt physically interact and regulate parathyroid hormone-dependent MMP13 expression in osteoblasts through a new osteoblast-specific element 2/AP-1 composite element". The Journal of Biological Chemistry. 276 (23): 20029–38. doi:10.1074/jbc.M010601200. PMID 11274169.
- ^ a b D'Alonzo RC, Selvamurugan N, Karsenty G, Partridge NC (January 2002). "Physical interaction of the activator protein-1 factors c-Fos and c-Jun with Cbfa1 for collagenase-3 promoter activation". The Journal of Biological Chemistry. 277 (1): 816–22. doi:10.1074/jbc.M107082200. PMID 11641401.
- ^ Chakraborty S, Sinha KK, Senyuk V, Nucifora G (August 2003). "SUV39H1 interacts with AML1 and abrogates AML1 transactivity. AML1 is methylated in vivo". Oncogene. 22 (34): 5229–37. doi:10.1038/sj.onc.1206600. PMID 12917624.
- ^ Levanon D, Goldstein RE, Bernstein Y, Tang H, Goldenberg D, Stifani S, Paroush Z, Groner Y (September 1998). "Transcriptional repression by AML1 and LEF-1 is mediated by the TLE/Groucho corepressors". Proceedings of the National Academy of Sciences of the United States of America. 95 (20): 11590–5. Bibcode:1998PNAS...9511590L. doi:10.1073/pnas.95.20.11590. PMC 21685. PMID 9751710.
- ^ Puccetti E, Obradovic D, Beissert T, Bianchini A, Washburn B, Chiaradonna F, Boehrer S, Hoelzer D, Ottmann OG, Pelicci PG, Nervi C, Ruthardt M (December 2002). "AML-associated translocation products block vitamin D(3)-induced differentiation by sequestering the vitamin D(3) receptor". Cancer Research. 62 (23): 7050–8. PMID 12460926.
Further reading[edit]
- Nucifora G, Rowley JD (July 1995). "AML1 and the 8;21 and 3;21 translocations in acute and chronic myeloid leukemia". Blood. 86 (1): 1–14. doi:10.1182/blood.V86.1.1.bloodjournal8611. PMID 7795214.
- Perry C, Eldor A, Soreq H (March 2002). "Runx1/AML1 in leukemia: disrupted association with diverse protein partners". Leukemia Research. 26 (3): 221–8. doi:10.1016/S0145-2126(01)00128-X. PMID 11792409.
- Imai O, Kurokawa M, Izutsu K, Hangaishi A, Maki K, Ogawa S, Chiba S, Mitani K, Hirai H (March 2002). "Mutational analyses of the AML1 gene in patients with myelodysplastic syndrome". Leukemia & Lymphoma. 43 (3): 617–21. doi:10.1080/10428190290012155. PMID 12002768. S2CID 45854670.
- Hart SM, Foroni L (December 2002). "Core binding factor genes and human leukemia". Haematologica. 87 (12): 1307–23. PMID 12495904.
- Michaud J, Scott HS, Escher R (2003). "AML1 interconnected pathways of leukemogenesis". Cancer Investigation. 21 (1): 105–36. doi:10.1081/CNV-120018821. PMID 12643014. S2CID 19586636.
- Ganly P, Walker LC, Morris CM (January 2004). "Familial mutations of the transcription factor RUNX1 (AML1, CBFA2) predispose to acute myeloid leukemia". Leukemia & Lymphoma. 45 (1): 1–10. doi:10.1080/1042819031000139611. PMID 15061191. S2CID 10770839.
- Yamada R, Tokuhiro S, Chang X, Yamamoto K (September 2004). "SLC22A4 and RUNX1: identification of RA susceptible genes". Journal of Molecular Medicine. 82 (9): 558–64. doi:10.1007/s00109-004-0547-y. PMID 15184985. S2CID 9156168.
- Harada H, Harada Y, Kimura A (September 2006). "Implications of somatic mutations in the AML1/RUNX1 gene in myelodysplastic syndrome (MDS): future molecular therapeutic directions for MDS". Current Cancer Drug Targets. 6 (6): 553–65. doi:10.2174/156800906778194595. PMID 17017876.
External links[edit]
- RUNX1+protein,+human at the U.S. National Library of Medicine Medical Subject Headings (MeSH)
- Overview of all the structural information available in the PDB for UniProt: Q01196 (Human Runt-related transcription factor 1) at the PDBe-KB.
- Overview of all the structural information available in the PDB for UniProt: Q03347 (Mouse Runt-related transcription factor 1) at the PDBe-KB.